Illuminating function in spatial proteomics
The Naveni® in situ proximity ligation technology pushes the boundaries of fluorescence-based in situ methodology, enabling researchers to get the maximum information from every analysis by visualizing and quantifying proteins, their interactions and modifications, in situ at the molecular level. An ability to detect even low abundant proteins ensures that proteins can be investigated without the need to overexpress or modify in any way the natural environment of the cell. Data generated contributes to a deeper understanding of the biological mechanisms within diseased or normal tissues, responses to therapeutic drugs or changes to the cellular microenvironment. The Naveni® in situ proximity ligation technology provides clean, highly resolved images, ready for quantification.
The benefits of our technology are:
- Clear target detection
- Consistency in staining
- Reproducibility
- Possibility to detect extremely low abundant proteins
- Detection of protein-protein interactions
- Specific detection of post-translational modifications, also with the use of pan-specific antibodies
Our technology is tailored for academic and industrial researchers working in fields of:
- Immuno-profiling
- Signaling profiling
- Drug testing
- Treatment validation
- Biomarker discovery
- Diagnostics
Workflow
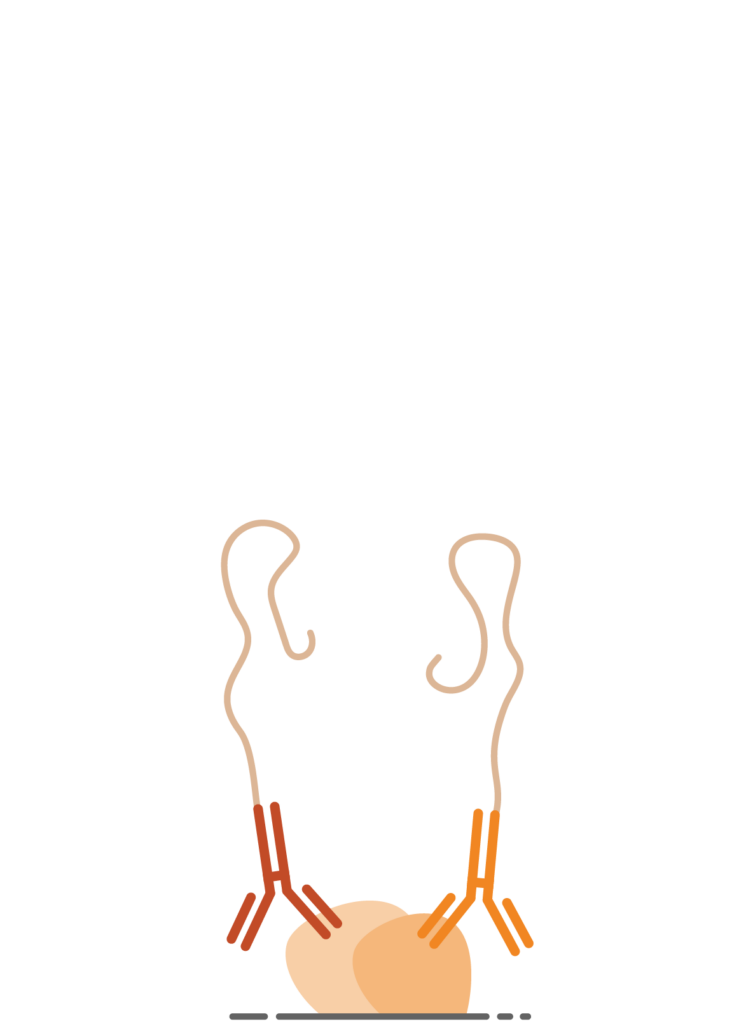


1. Proximal Binding of Navenibodies
The process begins when two Navenibodies are brought in close proximity by binding their target protein(s). This requirement for proximity by pairs of navenibodies is essential — it serves to enhance specificity and can indicate that the target proteins are interacting as parts of the same complex.
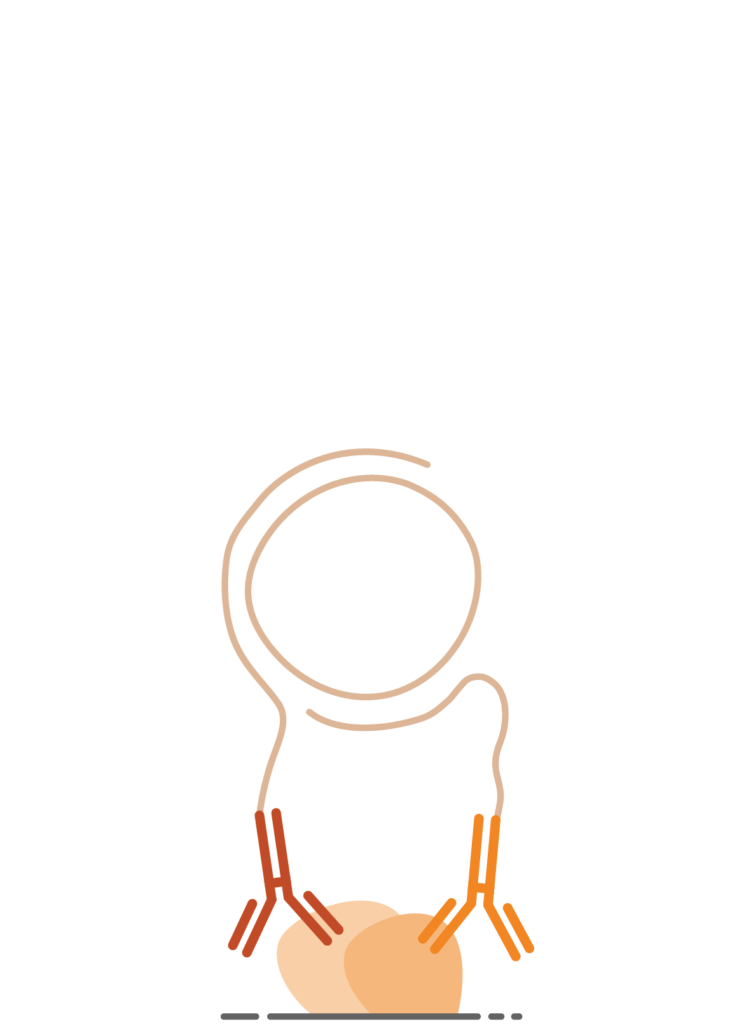
2. DNA Circle Formation
Navenibodies bound in close proximity guide the formation of a circular DNA template.
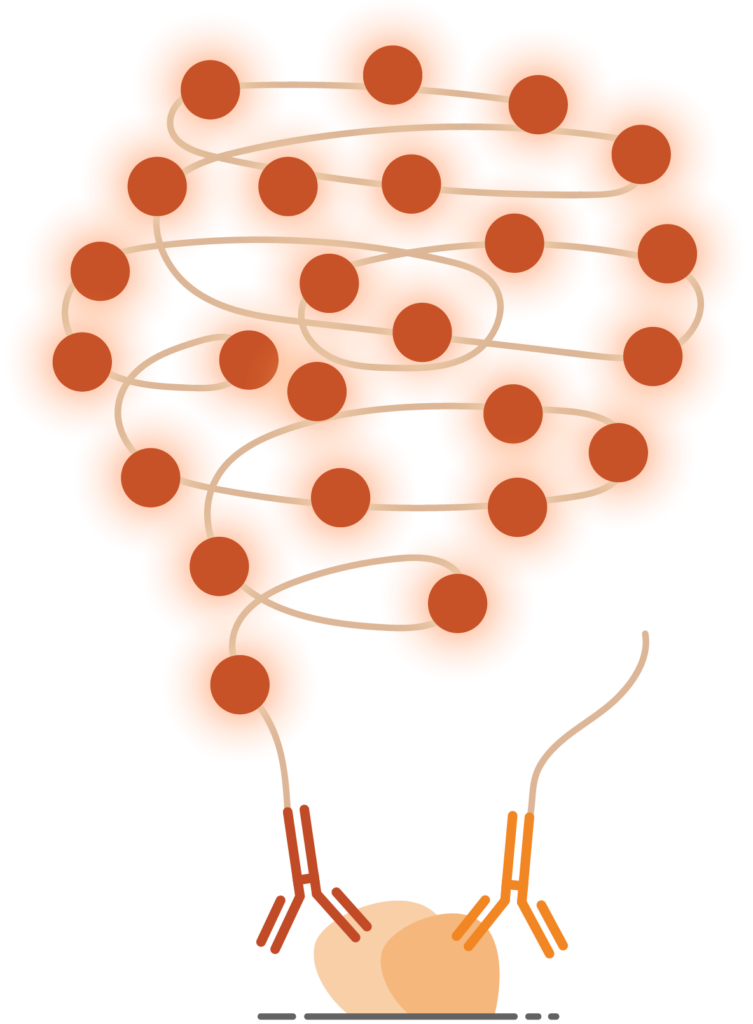
3. Amplified Signal
The circular DNA is amplified via rolling-circle amplification (RCA). The reaction products can be detected as intense, localized signals for highly sensitive detection reactions.
4. Data Visualization
The amplified signals are recorded under a microscope, visualized as distinct fluorescent dots at the site of the protein detection.
5. Data Quantification
These signals be quantified manually or using image analysis software, allowing researchers to monitor protein abundance and activity states in situ.
Workflow
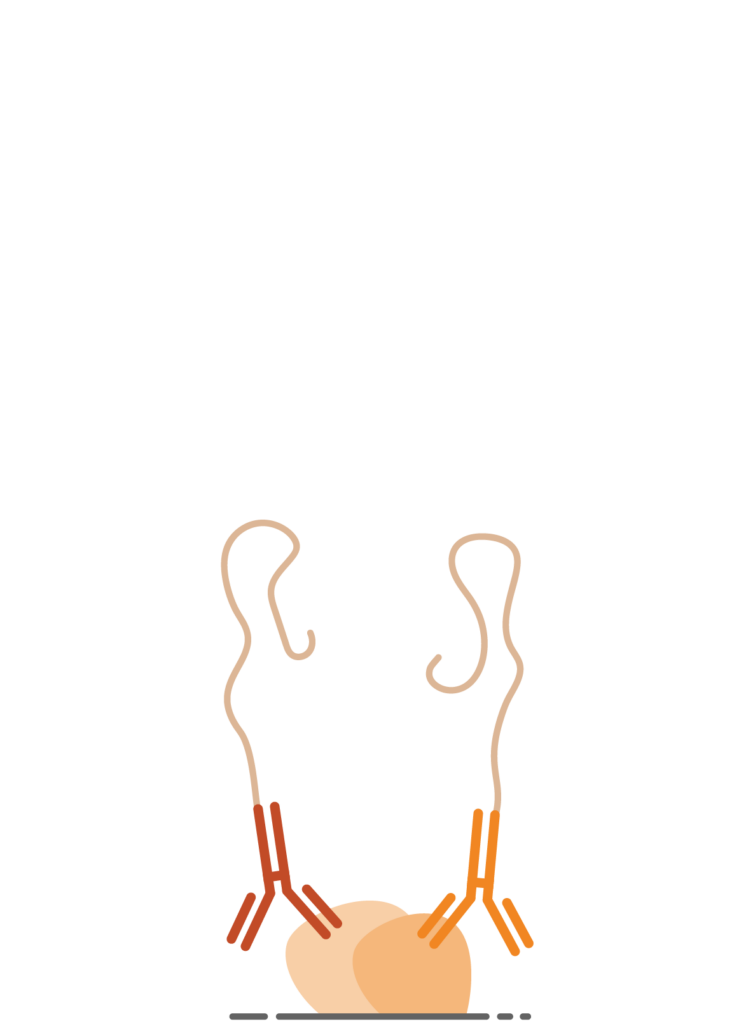
1. Proximal Binding of Navenibodies
The process begins when two Navenibodies are brought in close proximity by binding their target protein(s). This requirement for proximity by pairs of navenibodies is essential — it serves to enhance specificity and can indicate that the target proteins are interacting as parts of the same complex.

2. DNA Circle Formation
Navenibodies bound in close proximity guide the formation of a circular DNA template.

3. Amplified Signal
The circular DNA is amplified via rolling-circle amplification (RCA). The reaction products can be detected as intense, localized signals for highly sensitive detection reactions.

4. Data Visualization
The amplified signals are recorded under a microscope, visualized as distinct fluorescent dots at the site of the protein detection.

5. Data Quantification
These signals be quantified manually or using image analysis software, allowing researchers to monitor protein abundance and activity states in situ.