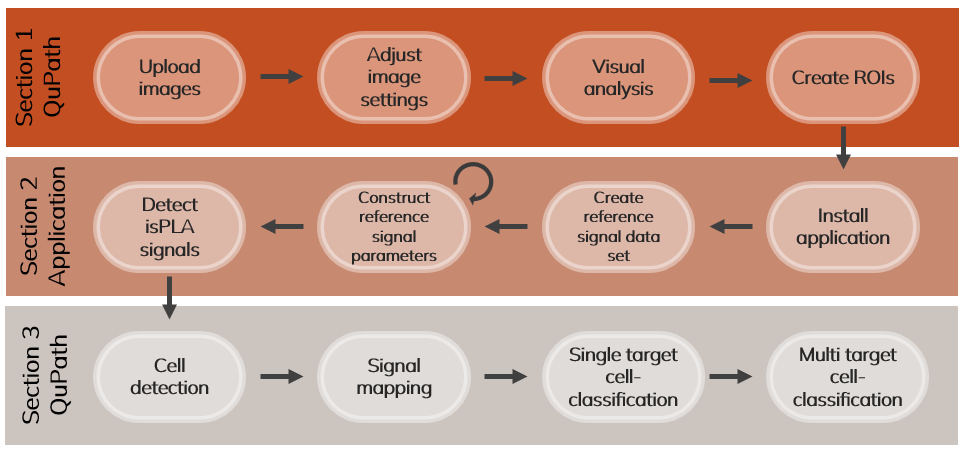
This guide aims to support QuPath users to analyse multiplex data from Navinci in situ proximity ligation assays (isPLA). The guide includes a brief introduction on how to use Qupath, annotate regions of interest (ROIs), how to perform isPLA signal detection, and examples of how to analyse the detected signals in QuPath by performing cell segmentation, mapping signals to cells by spatial distance and creating single or multi-target cell classification as shown in Figure 1.
QuPath Guidelines for Image Analysis
QuPath Guidelines for Image Analysis
Note on the demonstrated workflows:
IsPLA signal detection: QuPath offers limited options to detect and approximate the number of isPLA signals that are dense and form clusters. Therefore, the signal detection is executed in a separate application that communicates directly with QuPath to read images, annotations and import detection opjects. The spot detection functions are based on Big-FISH which uses gaussian mixtures to approximate the number of signals in clusters. For more information on Big-FISH, visit their documentation[1].
The remaining steps in this guide are intended as examples of how to analyse multiplex isPLA tissue images. Digital image analysis encompasses a wide variety of methods and strategies, which should be chosen based on the goal of the project. Cell segmentation is demonstrated with the StarDist extension due to its ease of use within Qupath and relatively good performance. However, evaluate which model is the best option for your analysis.
Requirements for this guide:
- Programs:
-
- Windows 11
- QuPath v0.5.1
-
The guide does not include:
- Recommendations for experimental set-up.
- Recommendations for visual assessment, tissue quality control and image- and data interpretation.
- Recommendations for pre-processing or image-normalization except for an example of background reduction when performing Big-FISH signal detection.
- Recommendations for downstream analysis.
1 Getting started with QuPath analysis
This section includes a brief introduction on how to create projects, add images and view images in QuPath, as shown in Figure 2. If you are comfortable operating QuPath, skip ahead to Section 2 for the signal detection/analysis. If you are completely new to QuPath, we recommend you review the comprehensive tutorials provided by the QuPath developers:
- Installation tutorials: found here
- A comprehensive guide of how to get started with QuPath (e.g. image viewing, creating annotations): found here
- QuPath image analysis tutorials: found here
- The QuPath paper: found here

1.1 Adding images to a project
1.1.1 Create a project by selecting the [Project] tab in the analysis menu and click on [Create project].
1.1.2 In the pop-up file explorer, create a new, empty folder in your desired directory and click on [Select Folder].
Note: A project is a good way to work with multiple images in QuPath. Within the project, one can easily switch between images, run batch-analysis and organize data files such as scripts and classifiers. Ensure to save edits and your current work before you close down QuPath or switch between projects and images. This is done by clicking on [File] → [Save as] or pressing Ctrl + s. To open an already existing project, select the [Project] tab and click on [Open project], select the project folder and then click on [Open].
1.1.3 To add images to the project, either drag and drop the images from the file explorer to QuPath or select the [Project] tab in the analysis pane menu and click on [Add images]. A dialogue box will pop up, as shown in Figure 3D. Click on [Choose files] and select your images in the file explorer and then click on [Open]. Select Fluorescence in the [Set image type] and then click on [import]. All images associated to the file will now be uploaded to the project.
Note: QuPath projects do not contain your images but has associated data files to locate them. If the images are moved to another location, a dialog box will pop up when you re-open the project with the option of re-defining the image locations. In the dialog box, click on [Search] and choose the correct directory, then click on [Apply Changes].
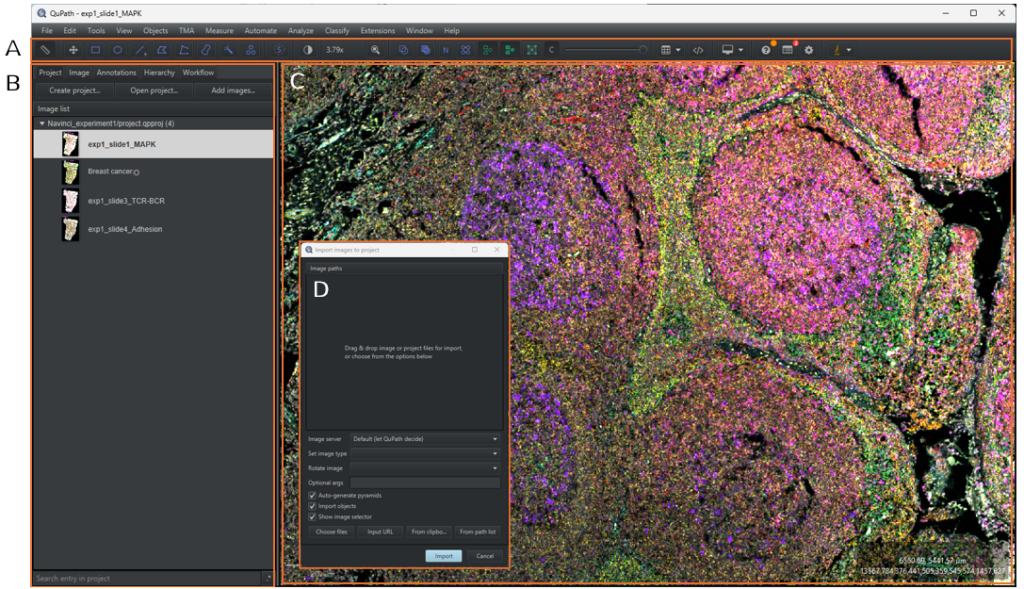
1.1.4 To open an image in the image viewer, double click on the image in the image list.
1.1.5 Go to the [Image] tab in the analysis menu to see the meta data of the image opened in the viewer. Information such as pixel size, image size, image URI and more can be found here.
1.1.6 Zoom in on the image by scrolling with the mouse or double-click on the Display magnification icon [y.yy] positioned to the left of the magnifying glass in the main menu. Go around in the image by dragging the mouse around in the image viewer.
1.2 Adjusting image settings
1.2.1 Click on [View] → [Show channel viewer] to open a window to see all the channels separately, as shown in Figure 4. By right clicking on the channel viewer the zoom can be set along with other settings. This is a helpful tool when adjusting image settings or when looking through detected and classified objects.
1.2.2 Click on the 
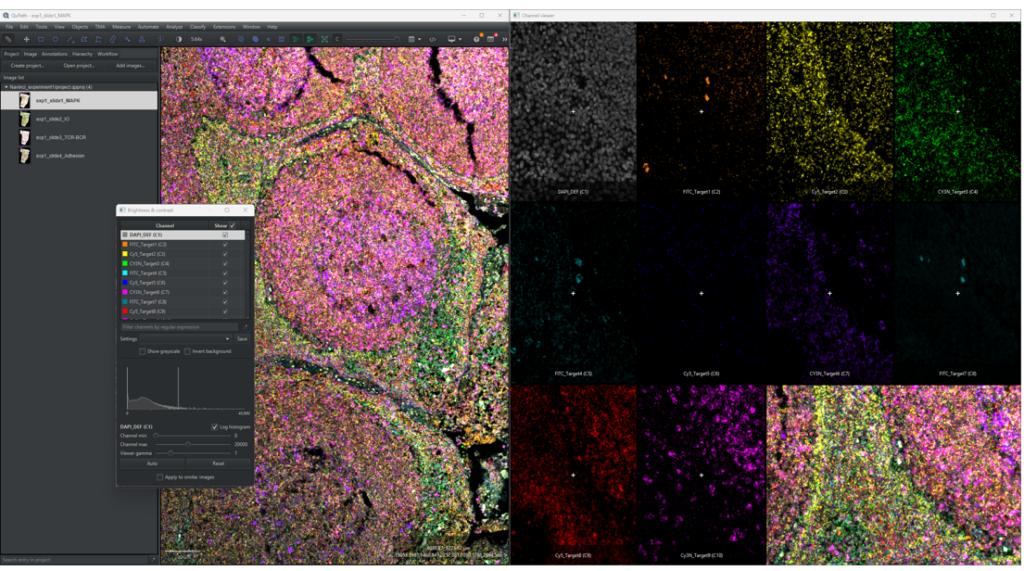
1.2.3 Go through each channel and change the [Channel max] value and [Channel min] value until you can clearly observe signals, if present, in each channel.
Note: Changing the [Channel min] will facilitate the overall interpretation of staining patterns, especially at high background and noise values. The minimum value should however be changed with caution since it can hide signals or overall cause difficulties in understanding what is signal and what is noise. It is good to go back to a minimum value at 0 to fully understand the signals at high magnifications at the individual channels.
1.2.4 (Optional) To change the names of the channels displayed in the Brightness & Contrast window to the names of the targets, you can double click on the channel names and type in a new name or use the code below to change all channels at once. Using target names instead of channel names facilitates data interpretation at later steps. To use the script:
-
-
- Open the script editor by clicking on [Automate] → [Script editor] or pressing ctrl + [, copy the code below and paste it into the editor.
-
-
-
- Add channels to the function by removing the
in rows 5-11 to activate them. For example, if you have six channels in total (with DAPI), then four additional rows should be added.
- Change the
in the active rows to the desired channel names. The code will name the channels by the order of the channel list, meaning that the first active row will name channel one (C1) and so on.
- Click on [Run] or press Ctrl + r.
- Add channels to the function by removing the
-
Note: It is very important that channel names are consistent in all images when running the scripts in this guideline, especially when creating and running the classifiers created in section 3.
1.2.5 The image settings can be saved by writing a file name in the [Settings] drop-down menu below the channel list and clicking on [Save] in the Brightness and contrast window. This enables the user to go back and forth between settings and apply the same settings between images (which may be easier than using the [Apply to similar images]). Note that this only works for images with the same channels, channel order and channel names.
1.3 Creating regions of interests
1.3.1 Click on one of the annotation drawing tools in the main menu to the left. In the ROIs shown in Figure 5 below, the rectangle tool, 

Tips and tricks when drawing regions of interests:
- To move annotations around, select the move tool by clicking on
or press m, and then select the annotation by double clicking on it. The annotation is moved by pressing the cursor down within the annotation and then dragging it around.
- To erase parts of the annotations, select the annotation by double clicking on it and select the brush tool,
. Hold down Alt and left click while moving the cursor around on the annotation.
- To rotate annotations, click on [Objects] → [Annotations] → [Transform Annotations] or press Ctrl + Shift + T. A small circle will appear above the annotation in the middle, click and hold on to it while dragging the mouse around to rotate. When you are happy with the results, push Enter and confirm changes by clicking on [Selected Object] in the pop-up window. Note that the transformation circle can be small and difficult to find.
- By clicking on [Objects] → [Annotations] in the main menu there are several nice-to-have tools for making regions of interests, such as [Create full image annotation], [Fill holes], [Expand Annotation] and much more.
1.3.2 Lock the annotation by right clicking on the object in the annotation list in the [Annotation] tab of the analysis menu and click on [Lock]. The setting prevents the annotation from being moved around.
1.3.3 When analysing different morphological regions in a tissue, it is easier to keep track of annotations by classifying them, as exemplified in Figure 5. Furthermore, the signal detection pipeline requires the annotations to be classified. To add a class to the classification list, go to the [Annotation] tab in the analysis menu and right click on the classification list located on the right of the Annotation list. Select [Add/Remove] → [Add class], write the class name in the pop-up window and click on [OK].
1.3.4 To classify a ROI, select the annotation in the annotation list, select the corresponding class in the classification list and click on [Set selected]. The annotation should now be outlined in the main viewer with the same colour as the class legend in the classification list and have the classification name written in parentheses next to its name.
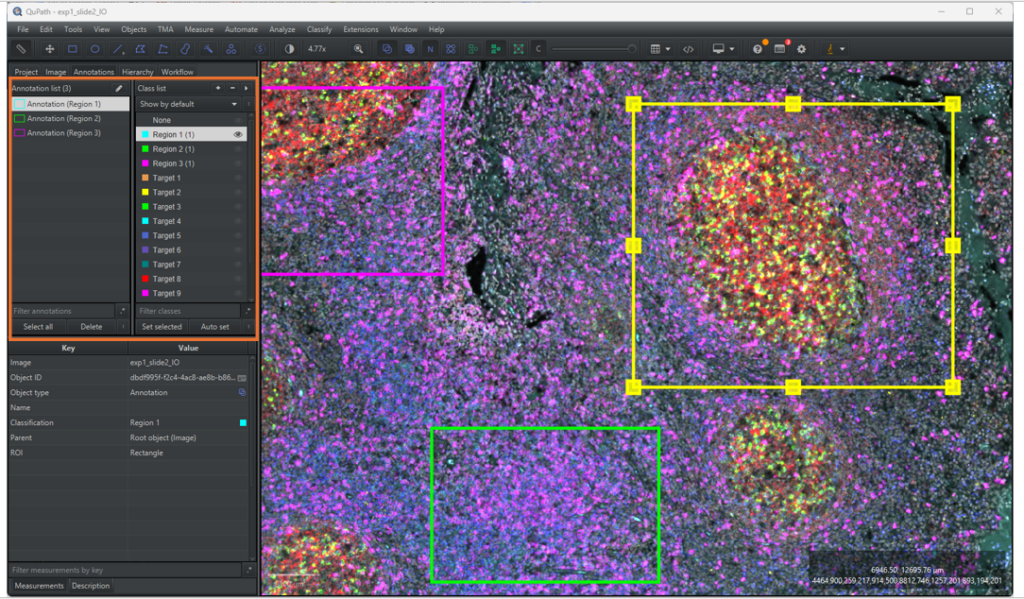
2 Signal detection
This section includes instructions and recommendations on how to detect isPLA signals. It is critical to understand isPLA signal features when optimising signal detection parameters. It is critical to understand isPLA signal features when optimising signal detection parameters as they can vary depending on the target (abundance and distribution), tissue and image acquisition settings. Signal features, as shown in Figure 6, to consider are:
-
- Sparse signals: Sparse signals can be in- or out-of-focus depending on where they are positioned relative to the z-plane that the image is taken in. Out-of-focus signals will have reduced intensity and an increase in apparent size. Accordingly, focused signals are easier to detect than out-of-focus signals.
- Clustered signals: Clusters consists of multiple signals with different x, y and z coordinates that are more or less visually indistinguishable from each other. Cell classification based on clustered signals can be performed by mean signal intensity.
- False positive signals: Although isPLA improves specific protein detection by requiring two binding events to form a signal, occasionally, false positive signals can still occur by proximal, unspecific antibody binding of probe pairs. False positives can be similar in shape and intensity compared to true positive signals but are often very sparse. Use of appropriate technical and biological controls can aid in distinguishing the spatial location and distribution of true and false signals.

Although sparse signals can be reliably detected using filtering and thresholding, these approaches are inadequate for resolving clustered signals. Gaussian mixture modelling addresses this challenge by fitting reference signal parameters within clustered regions to localize individual signals. Since this signal detection capability is not available in QuPath (< v0.5.1), it’s implemented in a separate application that communicates with QuPath, thereby extending its functionality.
This section will provide instructions for installation, parameter selection and parameter optimisation for signal detection. The application consists of three main functions: (1) sparse signal detection (Section 2.2), (2) reference signal construction (Section 2.3) where signal parameters are fitted based on a data set and (3) isPLA signal detection (Section 2.4) where both sparse signals and clustered signals are detected by the fitted signal parameters. For images mainly containing sparse signals, such as cell samples, we recommend following the instructions in Section 2.2•(function 1, sparse signal detection). For images containing clustered signals, we recommend to first follow the instructions in Section 2.2 (function 2, reference signal construction) and then follow the instructions in Section 2.3 (function 3, isPLA signal detection). Alternatively, follow section 2.3 first with a small ROI and example signal parameters, to understand the overall workflow, and go back to Section 2.2 to fit signal parameters.
The remaining sections contains useful information for setting up the application and parameter optimisation. Section 2.1 gives instructions for application installation and how to configure QuPath. Section 2.5 have instructions for how one can analyse the output signal detections in QuPath and Section 2.6 have recommendations for optimisation of signal detection parameters. For any errors and warnings printed by the application, see Section 2.7.

Note: For more advanced Python users, it is possible to interact with QuPath directly from Python via the library PAQUO3, which eliminates the need of importing and exporting annotation and detection objects. A script for using the library is given at the end of section 2.2. It is important to note that using the library requires calibrating the library with the QuPath software, which can be tricky for more novice Python users. If you would like to use PAQUO, you can skip section 2.1 and analyse your images with the script “PAQUO_signal_analysis.py” given at the end of section 2.3. Section 2.2 will help you modify all parameters correctly to run the analysis.
2.1 First time users
Please note that installation of the isPLA signal detection application requires Windows 11. Please contact our customer support for usage with another OS.
2.1.1 Installation
2.1.1.1 Download a zip with the installation file by clicking on the button below:
2.1.1.2 Extract the installer file “isPLA_signal_detection_installer” from the downloaded zip.
2.1.1.3 Initiate the installer by double clicking on the installer.exe file. Step through the installer until it is complete, make sure to agree to set up a shortcut on your desktop and start menu.
2.1.1.4 When the installation has run, the application will be initiated automatically and a dialogue window will appear on your screen, as shown in Figure 8 below. Note that it might take some time for the application to be initiated. If nothing appears, look for the icon in the taskbar.
2.1.2 QuPath configuration
When you open the signal detection application for the first time, QuPath might need to be configured before continuing with signal analysis. The configuration pane is visible at the top of the application window denoted “QuPath Configuration”. If a red cross and a text “Unsuccessful configuration” is shown in the pane, as shown to the right in Figure 8, then the application needs to be configured.
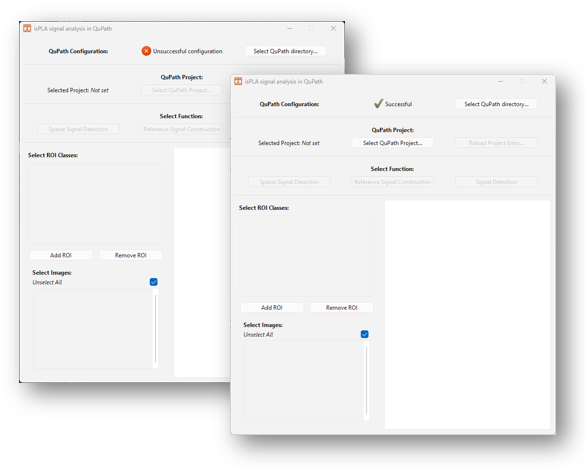
Configuration is done by the following instructions:
2.1.3 Click on the button [Select QuPath directory…].
2.1.4 In the pop-up directory, locate the file directory of your QuPath installation. If you don’t know where the application is stored on your computer, the file location can usually be found by right-clicking on a short-cut of the application and clicking on “Open file location”. If the file location points to a new short-cut, then right-click on the short-cut and select “Open file location”.
2.1.5 Select the QuPath-[version].exe file in the pop-up directory and click on [Open].
2.1.6 The configuration has worked when the red cross and text has been replaced with a green check mark and “Successful” text.
2.2 Sparse spot detection
This section provides instructions for how to detect sparse isPLA signals in images using the ‘isPLA signal detection’ application. Sparse signal detection can be used for when only sparse signals are present in the images or for when creating a reference data set for building signal parameters, as further instructed in Section 2.3.
Note that steps 2.2.1 and/or 2.2.2 can be skipped if the ‘isPLA signal detection’ application has already been initialized and/or a project has already been opened. If there have been any new changes to the project while open in the application, make sure to reload the project by clicking on the button [Reload Project Entry…].
2.2.1 Initialize the ‘isPLA signal detection’ application by double-clicking on the desktop short-cut, alternatively search for “isPLA signal detection” on your computer.
2.2.2 Click on the button [Select QuPath Project…] and locate your QuPath project file (‘project.qpproj) in the directory and click on ‘Open. Wait until the ‘Selected Project:’ has changed from ‘Not set’ to the name of the QuPath project parent folder and the box ‘Select images:’ have been updated with the images in your project.
2.2.3 Under “Select Function”, click on the button [Sparse Signal Detection], as shown in Figure 9.
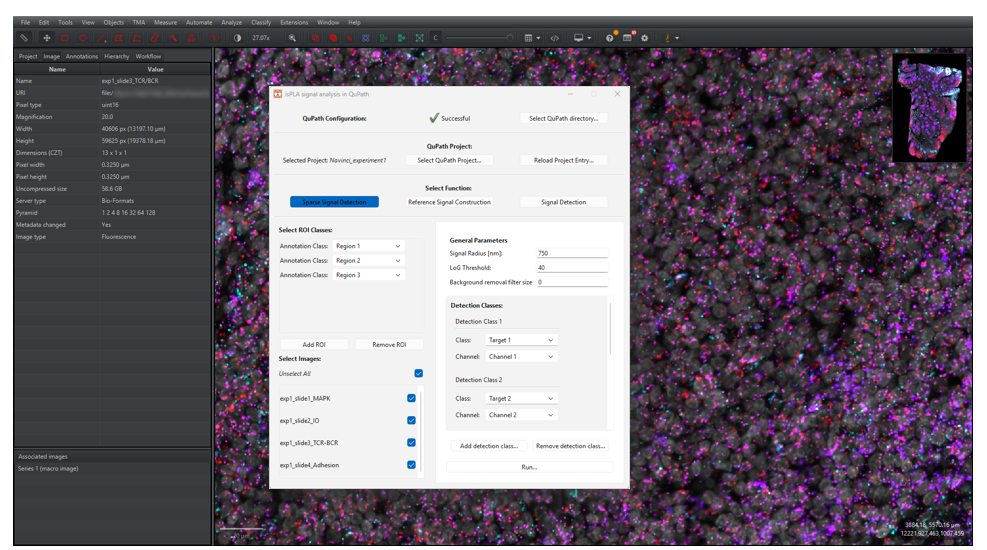
2.2.4 Select ROI classes under the “Select ROI classes” on the left side of the menu:
- If you have created ROI annotations, as instructed in Section 1.3, then click on [Add ROI] on the left side of the window. Select the class in the dropdown menu “Annotation class”. If the class is not available, the changes might not have been saved to QuPath properly before opening the project in the application. Make sure to resave and then reload the project in the application by clicking on [Reload Project Entry…].
- To add more classifications, click on [Add ROI] again, and to remove classifications click on [Remove ROI].
- To analyse the whole images, make sure that no classifications are added.
2.2.5 Select or deselect images to analyse under “Select images” on the left hand-side.
Note that if you make any changes to the QuPath project, such as creating new ROIs or adding new images, while having the project open in the application, make sure that your QuPath project is saved and click on the button [Reload Project Entry…] in the application.
2.2.6 Go to the “General Parameters” on the right side of the window. Either leave the default values as is for now or read about the parameters below.
Spot radius: select a value that reflects the expected signal radius in nano meters. You can measure distances in QuPath by drawing a line using the line annotation tool and checking the “Length µm” measurement.
Log Threshold: the parameter decides the threshold for detecting signals in the Laplacian of Gaussian filtered image. Since the filter enhances circular objects of a specific size, a greater threshold will result in a stricter detection regarding size and circularity. Keep in mind that higher intensity objects will have a greater value compared to lower intensity objects.
Background removal filter size: the parameter decides the filter size for the background removal. Setting the parameter to zero will make the application skip this step entirely. The filter operates by calculating the mean intensity value within a neighbourhood around each pixel and subtracting the mean from the pixel’s raw intensity. The size of the neighbourhood is defined by the parameter, e.g. a value of 10 will calculate the mean in a 10×10-pixel region. Use this setting with care: the mean value can be strongly influenced by unrepresentative high intensity pixels, and regions with varying signal densities are affected differently. In addition, larger filter sizes require more computation time.
2.2.7 Under “Detection classes”, select the desired output of the signal detection:
- The “class” refers to the class name that the detections will be exported to QuPath with.
- The “Channel” refers to the channel index, starting from zero, in which the detection should occur.
- Detection classes can be added or removed by clicking on [Add Detection Class…] and [Remove Detection Class…].
For the example shown in Figure 9, signals will be detected in channels 1 and 2 and exported as classes “Target 1” and “Target 2” respectively.
Note that new classes can be added by clicking on the dropdown menu and typing a new name.
2.2.8 Click on [Run…] to start the analysis.
Note that it is not recommended to make changes to the QuPath project while the function is running, since it will cause either the application or QuPath to overwrite the changes or signal detection exports.
Note that dialogue windows will be raised, as shown in Figure 10, if:
- Parameters are missing
- Duplicate classes have been selected
- A large number of images have been selected
- No ROI classes are selected
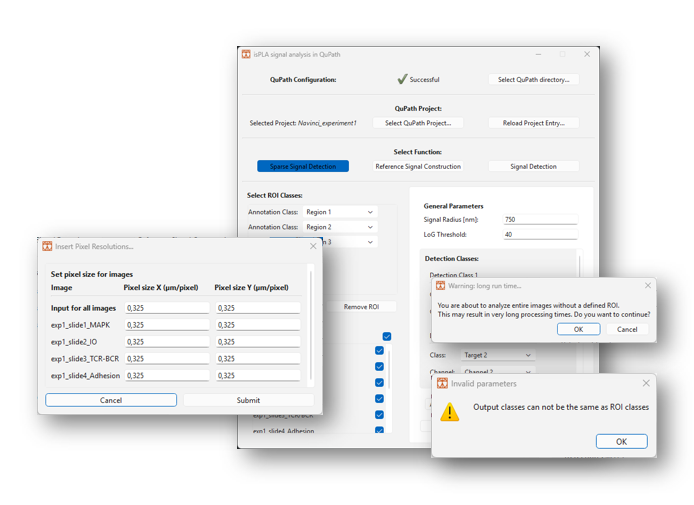
2.2.9 A new window will appear to update the user about the process, as shown in Figure 11. Once finished, the window will write out “Finished sparse spot detection” and “Saved edits to QuPath”. Click on the “Close button” to return.
Note that the application will save QuPath edits every five images, meaning that the print “Saved edits to QuPath” does not alone mean that the pipeline is finished.
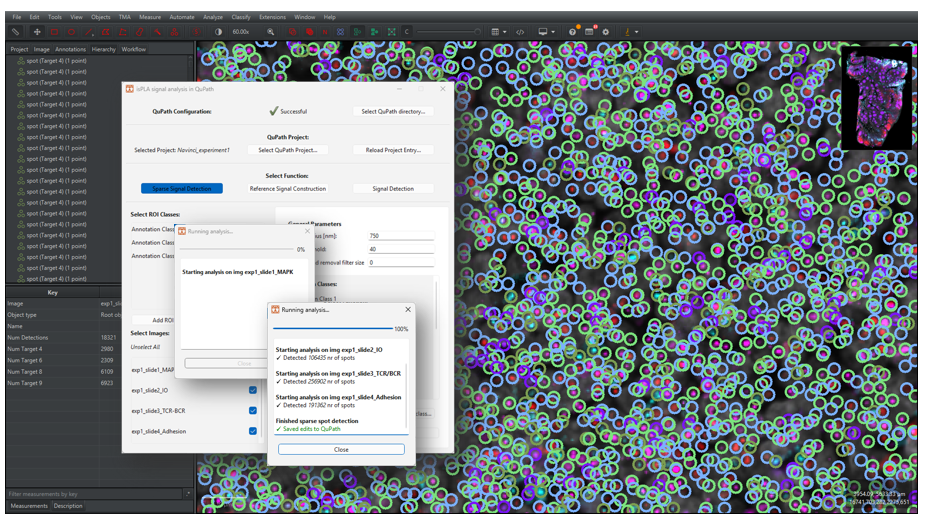
Note that any errors occurring during detection will be printed out in the window in red text, go to Section 2.7 for more information.
2.2.10 Go back to QuPath. If you have any images opened, make sure to click on [File] → [Reload Data] or press Ctrl + R. For instructions of how to visualize signals in QuPath, go to Section 2.5.
Note, to clean up the hierarchy list, click on [Objects] → [Annotations] → [Resolve hierarchy]. By doing so, the detection objects will be assigned as child objects to any overlapping annotation.
2.3 Fit signal parameters to a reference data set
This section provides instructions for how to fit signal parameters for isPLA spot detection. Please note that signal parameters depend on the filter and exposure time used during image acquisition.
2.3.1 Annotate image regions containing sparse isPLA signals in your QuPath project.
Note that steps 2.3.2 and 2.3.3 can be skipped if the ‘isPLA signal detection’ application has already been initialized and/or a project has already been opened. If there have been any new changes to the project while open in the application, make sure to reload the project by clicking on the button [Reload Project Entry…].
2.3.2 If the application is not already opened, initialize the ‘isPLA signal detection’ application by double-clicking on the desktop short-cut, alternatively search for “isPLA signal detection” on your computer.
2.3.3 Click on the button [Select QuPath Project…] and locate your QuPath project file (‘project.qpproj) in the directory and click on ‘Open. Wait until the ‘Selected Project:’ has changed from ‘Not set’ to the name of the QuPath project parent folder and the box ‘Select images:’ have been updated with the images in your project
2.3.4 Run the sparse signal detection function within the annotated sparse signal regions by following the instructions in Section 2.2.
Note that several classes should be added to fit signal parameters between channels and or filter sets.
Optionally, one can go through the resulting detections in QuPath, making sure that annotated signals are well-differentiable from other signals and that there are no false positive annotations.
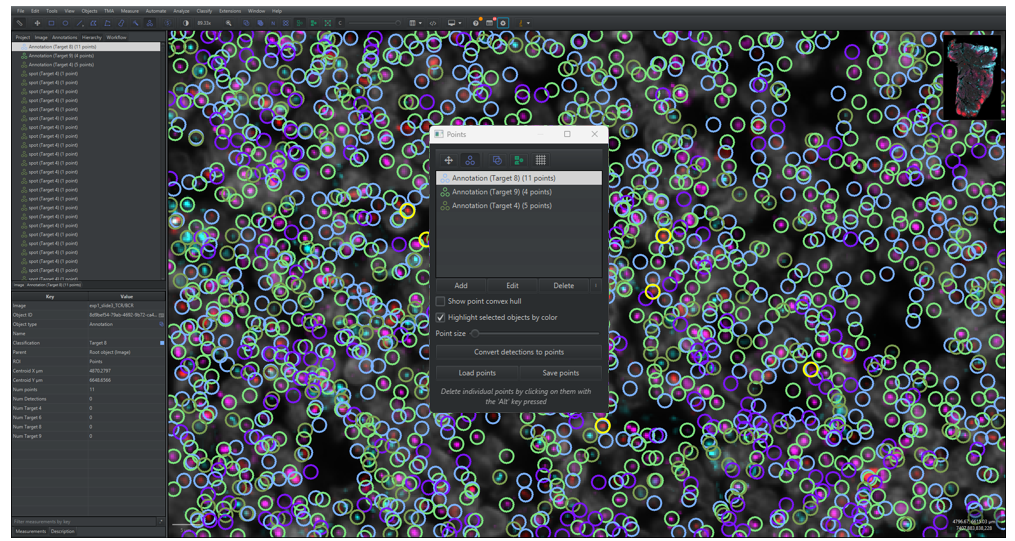
2.3.5 Optionally go through the resulting detections in QuPath and remove and add signal annotations, as shown in Figure 12. Annotated signals should be well-differentiable from other signals.
- Remove false positive signals by selecting them and clicking on [Objects] → [Delete…] → [Delete selected objects] or by pressing Delete. Press down Ctrl to enable selection of multiple point objects.
- Add points by…
- Open the Points dialogue window by clicking on the Points symbol,
, in the main menu, or by pressing “.” on your keyboard.
- Click on [Add] in the dialogue window to add a new class of points. Select the correct class in the Class list in the analysis pane in the [Annotations] tab and click on [Set Selected].
- To add points, make sure that the correct points group is selected and that the points symbol is marked and then click on a signal on the image. Make sure that the point is centered on the signal.
- Open the Points dialogue window by clicking on the Points symbol,
2.3.6 Make sure to save the changes in QuPath before continuing by clicking on [File] → [Save] or by pressing Ctrl S
2.3.7 Go back to the ‘isPLA signal detection’ application. Make sure to reload the project by pressing [Reload Project Entry”]. Click [OK] on the pop-up window.
2.3.8 Click on [Reference Signal Construction] under “Select Function”, as shown in Figure 13 below.
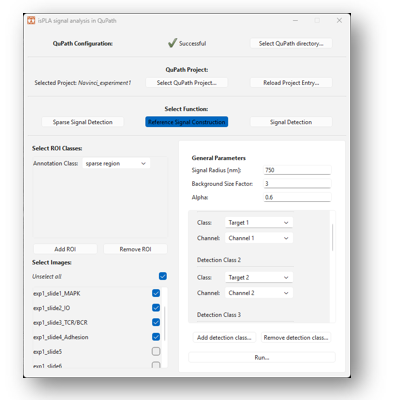
2.3.9 Optionally add ROIs under the “Select ROI classes”. This is done by clicking on [Add ROI] under the [Select ROI classes] on the left side of the window. Select the classification in the dropdown menu “Annotation class”. To add more classifications, click on [Add ROI] again, and to remove classifications click on [Remove ROI].
2.3.10 Select or deselect images to analyse under “Select images” on the left hand-side.
2.3.11 Go to general parameters on the left side of the window.
Spot radius: select a value that reflects the expected signal radius in nano meters. You can measure distances in QuPath by drawing a line using the line annotation tool and checking the “Length µm” measurement.
Background size factor: specifies the ratio between the minimum size of background objects and the size of the actual signal. The ratio is used to remove background in the reference data set images to get a better approximation for the signal parameters. A value of zero means that the preprocessing background removal is skipped.
Alpha: The parameter ‘Alpha’ refers to the intensity percentile used to compute the reference spot. For example, setting alpha to 0.5 results in building a mean signal from the upper 50% quantile of all spots. With other words, a greater alpha, brighter spots will be used to build a mean signal.
2.3.12 Select channels and output classes in the section “Detection classes”. This is done by selecting the input detection classes and their corresponding channel index for which point objects have been saved in QuPath.
For the example shown in Figure 12, points have been added in classes “Target 8” for channel index 8, meaning that the class “Target 8” with channel index 8 should be added as a detection class. Detection classes can be added or removed by clicking on [Add Detection Class…] and [Remove Detection Class…].
2.3.13 Click on [Run…] to start the analysis. A new window will appear to update the user about the process, as shown in Figure 14.
2.3.14 Once finished, the window will write out “Finished building reference parameters. Saved parameters to file …”. Click on the “Close” button to return.
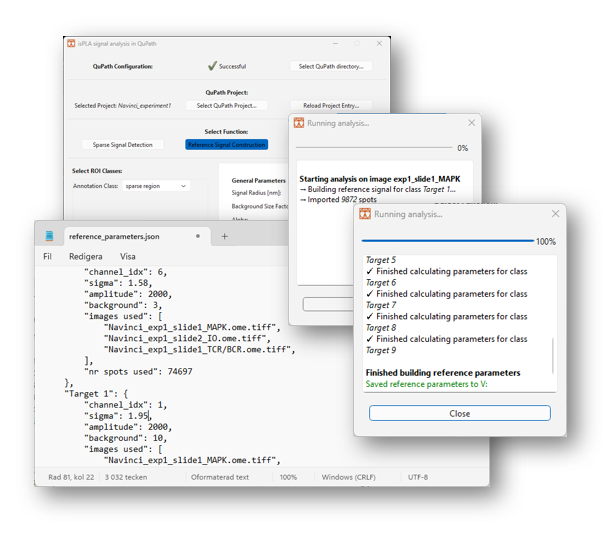
The resulting signal parameters will be saved in json files in the directory of your QuPath project. The file “reference_parameters.json”, as show in Figure 14, contains mean parameter values from all images selected in the analysis, and the files “reference_parameters-[image name].json” contains values from the individual images.
- The parameters are written as a dictionary where each detection class is a key and their value is a dictionary with the specific parameters belonging to that class, which are described below.
- “channel_idx”: the channel index (starting from zero) that has been used to build the parameters
- “sigma”: the signal radius in pixels
- “amplitude”: the signal intensity at its peak
- “background”: the intensity of the background surrounding the signals
- “images used”: a list with all images used to build the reference signal
- “nr spots used”: the number of spots used to build the reference signal
Note, if you want to use the same signal parameters for multiple detection classes when running “isPLA spot detection”, then you can open the reference parameters file and copy and paste the reference signal detection class and change the “channel_idx” and class name accordingly.
2.3.15 Reload your QuPath project in the application by pressing the button [Reload Project Entry…] and press [OK] on the pop-up dialogue.
2.3.16 When the project is reloaded, the application will search for a reference parameter file within the QuPath project folder. If the file is found, the detection classes in the function “isPLA signal detection” will be updated. Click on [Signal Detection] to see if the detection classes have been updated.
2.4 isPLA signal detection
This section provides instructions for how detect both sparse and clustered signals.
Note that steps 2.4.1 and 2.4.2 can be skipped if the ‘isPLA signal detection’ application has already been initialized and/or a project has already been opened. If there have been any new changes to the project while open in the application, make sure to reload the project by clicking on the button [Reload Project Entry…].
2.4.1 If the application is not already opened, initialize the ‘isPLA signal detection’ application by double-clicking on the desktop short-cut, alternatively search for “isPLA signal detection” on your computer.
2.4.2 Click on the button [Select QuPath Project…] and locate your QuPath project file (‘project.qpproj) in the directory and click on ‘Open. Wait until the ‘Selected Project:’ has changed from ‘Not set’ to the name of the QuPath project parent folder and the box ‘Select images:’ have been updated with the images in your project.
2.4.3 Under “Select Function”, click on the button [Signal Detection], as shown in Figure 15 below.
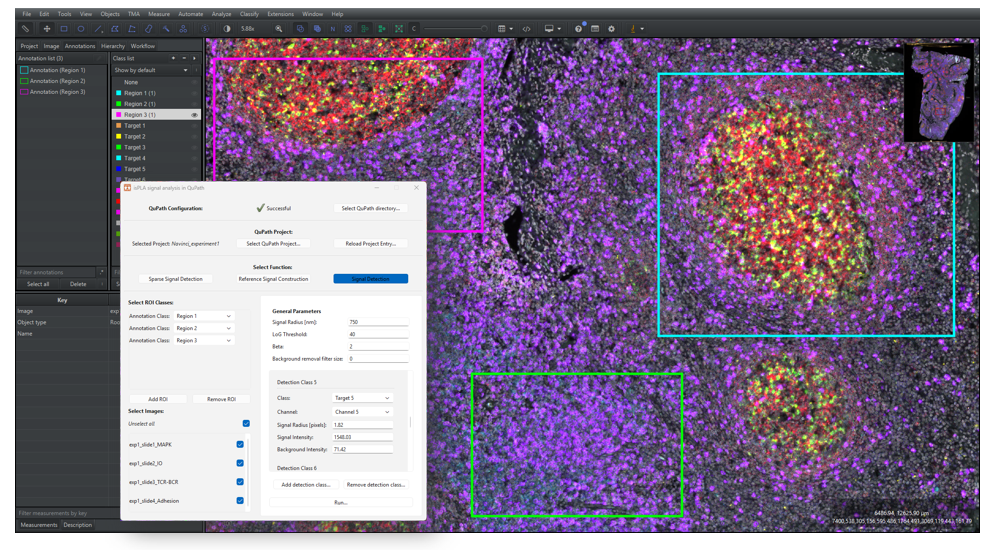
2.4.4 If you have created ROI annotations, as instructed in Section 1.3, then click on [Add ROI] under the [Select ROI classes] on the left side of the window. Select the classification in the dropdown menu “Annotation Class”. To add more classifications, click on [Add ROI] again, and to remove classifications click on [Remove ROI]. To analyse the whole images, make sure that no classifications are added.
2.4.5 Select or deselect images to analyse under “Select images” on the left hand-side.
Note that if you make any changes to the QuPath project, such as creating new ROIs or adding new images, that should be updated in the application, make sure that your QuPath project is saved and click on the button [Reload Project Entry…] in the application.
2.4.6 Go to the “General Parameters” on the right side of the window. Either leave the default values as is for now or read about the parameters below.
Spot radius: select a value that reflects the expected signal radius in nano meters. You can measure distances in QuPath by drawing a line using the line annotation tool and checking the “Length µm” measurement.
Log Threshold: the parameter decides the threshold for detecting signals in the Laplacian of Gaussian filtered image. Since the filter enhances circular objects of a specific size, a greater threshold will result in a stricter detection regarding size and circularity. Keep in mind that higher intensity objects will have a greater value compared to lower intensity objects.
Beta: the parameter is a factor used in the threshold for detection of clustered regions. In short, a greater beta means that only brighter regions will undergo cluster decomposition. In detail, a clustered region is every region of at least two pixels that have an intensity above the expected signal intensity (see Detection classes) multiplied by beta.
Background removal filter size: the parameter decides the filter size for the background removal. Setting the parameter to zero will make the application skip this step entirely. The filter operates by calculating the mean intensity value within a neighbourhood around each pixel and subtracting the mean from the pixel’s raw intensity. The size of the neighbourhood is defined by the parameter, e.g. a value of 10 will calculate the mean in a 10×10-pixel region. Use this setting with care: the mean value can be strongly influenced by unrepresentative high intensity pixels, and regions with varying signal densities are affected differently. In addition, larger filter sizes require more computation time.
2.4.7 Under “Detection classes”, select the desired output of the signal detection:
If a reference signal has been constructed for the project previously, this section should be filled in automatically. Classes can be removed or added by using the [Add detection class…] and [Remove detection class…] buttons. Note that the detection class list can be reset by reloading the project, although it also resets all other parameter selection.
If a reference signal has not been constructed for the project previously, then add parameters to each detection class accordingly:
- “Classification”: refers to the output classification exported to QuPath. Note that a new class can be added by clicking on the drop-down menu and typing in the name.
- “Channel” refers to the channel index, starting from zero, in which detection should occur.
- “Signal Intensity” refers to the expected signal intensity at its peak. The parameter depends both on image bit-depth, filter set and exposure time. To find a good starting value, one can hover on a signal in QuPath and read the intensity values in the lower left corner.
- “Signal radius [pixels]” refers to the expected signal radius in pixels. Signals are expected to have a radius around 500-750nm.
- “Background Intensity” refers to the expected background intensities. This can be set to zero if the images have been pre-processed beforehand to remove background.
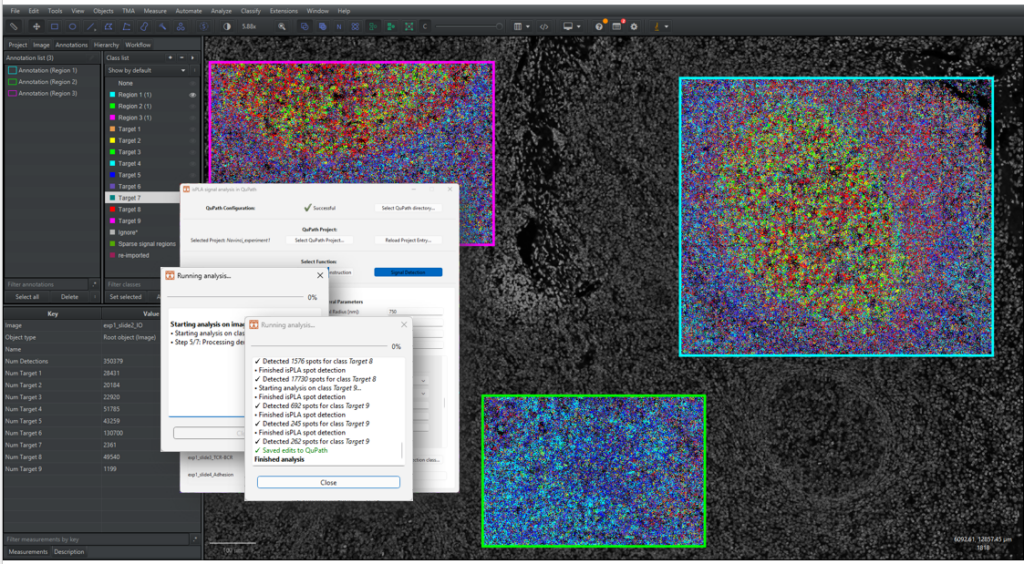
2.4.8 Click on [Run…] to start the signal detection. A new window will appear to update the user about the process, as shown in Figure 16 below. Once the process is finished, the window will write out “Finished Analysis”. Click on the “Close” button to return.
Note that it is not recommended to make changes to the QuPath project while the function is running, since it might cause QuPath the overwrite the changes.
Note that detections are exported and saved to QuPath for each image. The file “logbook.txt” saves each run, parameters and for which images the run has been successful. If the application would quit unexpectedly, please check the logbook to see which images were run successfully.
- Go back to QuPath. If you have any images opened, make sure to click on [File] → [Reload Data] or press Ctrl + R. For instructions of how to visualize signals in QuPath, go to Section 2.5.
2.5 Visualizing signals on QuPath
2.5.1 Open one of the analysed images in your QuPath project. The signal detection should now be visible in the main viewer, as shown in Figure 17 below. If they are not visible, try clicking on 
2.5.2 Go to the [Hierarchy] tab in the analysis pane to see a list of the detection signals. The signals will be listed as “(Type) (Target X) (1 Point)”, where “Type” is either a “Spot” or a “Clustered spot”.
2.5.3 The detected signals are saved as Points, meaning that their geometry only consists of a coordinate. To visualize the points better, QuPath shows a circle surrounding the coordinate. The size of the circles can be changed by clicking on the symbol 
Note: Changing the point size can be slow for a large amount of data.
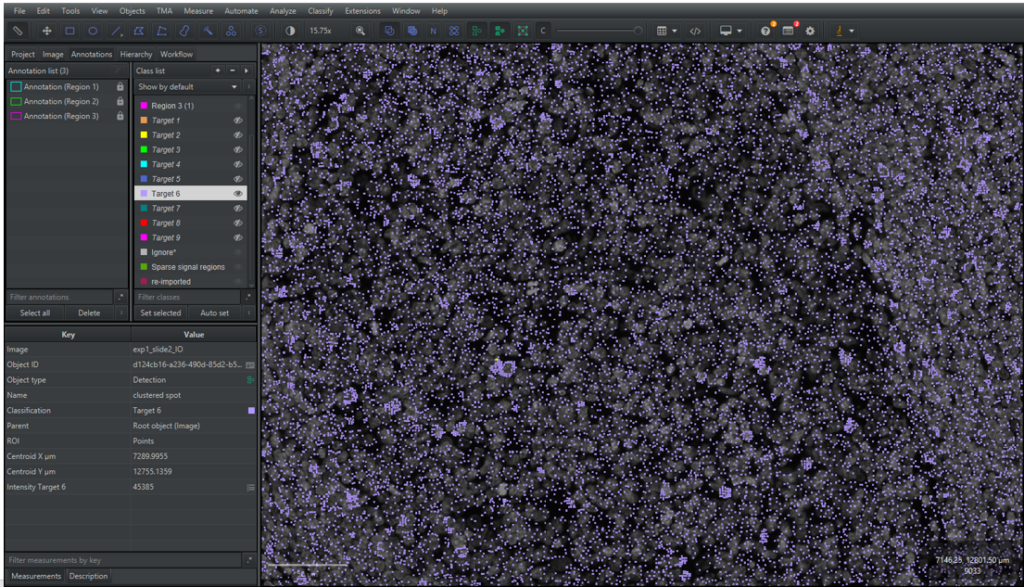
2.5.4 Unselect the signal and go to the [Annotation] tab in the analysis menu. At the bottom of the analysis pane there is a summarisation of the number of detections in the image for each class.
2.5.5 To modify the appearance of signal detections in the viewer, the classes need to be added to the Classification list. Go to the [Annotation] tab in the analysis menu and right click in the Classification list, click on [Populate from existing objects] → [Base classes only] and click [Yes] in the pop-up window.
2.5.6 Go through the detection classes one by one by toggling off and on the channels and the overlay view of the classes by right clicking on the class in the classification list → [Show/Hide…] → [Hide classes in viewer] / [Show classes in viewer] or by selecting the class/classes and pressing h / s. Use the channel viewer, as explained in Section 1.2, to look at the channels separately.
2.5.7 Review the signal detection in the ROIs. If you are not satisfied with the results, see section 2.6 below for tips on how to change parameters and re-do the signal detection.
2.6 Signal detection optimization
This section provides some guidance regarding the most common issues in signal detection and what parameters to adjust to alleviate them. Changing the signal detection requires deletion of previous detections as all signals must be re-detected with new parameters. Instructions for how to delete detections and re-run signal detection are found at the end of this section. Make sure to read through the optimization explanations below before deleting any detection objects.
Too many false negative sparse signals: There are non-detected signals in the image
- The LoG threshold used in the signal detection may be too high, try lowering it.
- If there are a lot of out-of-focus signals (blurry: low intensity and larger than in focus signals), the algorithm may not capture all signals. In that case, it is better to select a LoG threshold based on in-focus signals.
Too many false positive signals: Noise is detected as signals
- If the false positive signals are named “spot”, then the LoG threshold might be too low. Try to increase the threshold.
- If autofluorescent noise is detected as signals it is best to pre-process the images to remove background.
Decomposition of clustered regions are not as expected:
- Note that you can’t know how many signals there are in clustered regions, so be careful when changing signal decomposition parameters.
- The beta parameter is the multiplication factor of thresholding for a clustered signal region threshold = beta · amplitude). The parameter can therefore be changed to optimise cluster region detection. A greater beta means that brighter regions will be detected as clustered regions, resulting in generally a lower number of detected signals. A lower beta will on the other hand generally increase the number of detected signals.
- The gaussian parameter background can be changed to match the estimated background of your image. The parameter is important to not overestimate the number of signals in clusters.
- The background should preferably be measured in autofluorescent images (without fluorophores) with the same filters and exposure times.
- If you are not happy with your results, gaussian parameters can be fitted to your parameters by following the instructions in section 2.3.
Please do the following to re-run signal detection:
2.6.1 Delete the signal detections by their classification by right-clicking on their class name in the classification list in the [Annotation] tab in the analysis pane. Click on [Select objects by Classification] and all detections in the class will be selected. Click on [Objects] in the main menu → [Delete] → [Delete selected objects]. Repeat this for all targets you are dissatisfied with.
2.6.2 Reload the project in the “isPLA signal detection” application by clicking on [Reload Project Entry…]
2.6.3 Run the function with new parameters based on the explanations found in this section.
2.7 Application errors and warnings
This section gives explanations for the different types of errors and warnings that can be printed out in the application when running a function. Warnings are printed in orange, meaning that the script has completed the tasks. Errors are printed in red which means that the script could not complete the task for the image(s) and or detection class(es).
General warnings and errors:
- “Failed reading image”: The image is not readable by the application
- “Image is not readable”: The file URL cannot be found. This can be due to creating images within QuPath (for example by creating a training image).
- “No ROI annotations found in the image”: No annotations of the specified ROI class were found in the image. Make sure that the project is saved correctly and reload the project in the application. Note that by not specifying any ROI classes, the whole image will be analyzed
- “Out of memory”: No more memory can be allocated for the function.
Reference for signal construction warnings and errors:
- “Warning: low intensity mean signal”: The mean signal built from the reference data set has a low maximum intensity. If this warning has been printed, it is recommended to check the “amplitude” parameter in the output file. If the parameter is low compared to the maximum intensity of annotated signals in the reference data set, then the background size factor might be too low. Try to run the function without background removal by setting the factor to zero.
- “No signals found for class [class]”: No signals were imported from QuPath for the specified class
- “Modeling signals was not successful”: An error occurred when modeling the signal parameters.
- “Only [x] number of spots were used for class [class]”: A low number of reference spots were imported from QuPath.
- “No signal parameters found for class [class]”: This error could be due to multiple reasons and is due to the script not finding any signal parameters for the class.
- “Reference parameter export failed”: The export of the signal parameters could not save properly.
For warnings not explained here, please contact Navinci’s customer support. For quicker feedback, attach the file “log_errors.txt” found in the folder “_internal/isPLA_signal_detection/GUI” in the directory of the downloaded application.
3 Analysing signals in QuPath
This section demonstrates how signals detected with Big-FISH can be analysed in QuPath to classify cells. Figure 18 below provides an overview of the steps included.

The cell detection in this guideline is done with the StarDist extension, which segments stained nuclei, and the signals are mapped to cells using a distance-based approach. Per-target-classifiers are built by thresholding cells by the number of signals and are used to build a multi target cell classifier which classifies cells as, e.g. single and double positive. Lastly, this section will provide an example of how the results can be summarized in QuPath.
3.1 Cell detection by StarDist-QuPath extension
There are different possibilities of doing computational image analysis of multi-plex isPLA tissue images. One step that usually but not necessarily is included is cell detection, which enables the user to assign signals to cells and perform cell classification and phenotyping. In this section, StarDist cell detection in QuPath is exemplified. StarDist is a deep-learning-based nucleus detection method that can be used as an alternative method of cell detection in QuPath other than CellPose and QuPath’s own cell detection algorithm. It is quite straight-forward to use the extension, although the method requires scripting.
Running StarDist for the first time:
3.1.1 Go to the QuPath-Stardist-extension webpage[2] and download the latest file of qupath-extension-stardist-[version].jar from releases, make sure that the version is compatible with your version of QuPath. For example, for QuPath 0.5.0, download “qupath-extension-stardist-0.5.0.jar”.
3.1.2 Drag the stardist extension file into the main window of QuPath. You might have to restart QuPath before being able to access the extension.
3.1.3 Download a pre-trained StarDist model that is compatible with QuPath. In this guide, we use used “dbs2018_heavy_augment.pb”[3].
Running the StarDist extension in QuPath:
3.1.4 Open the script editor by clicking on [Automate] → [Script] editor or pressing ctrl + [. Copy and paste the code below in the editor, as shown in Figure 19. The script segments cells in the image by the StarDist extension with the use of a pre-trained model.
3.1.5 Change the path direction at row 22 to the path of the pre-trained model, in this case “dsb_2018_heavy_augment.pb”. Any “\” symbols in the path should be replaced with “/” or “\\”.
3.1.6 Change the ROI class names at row 19 so that they correspond to the classes of your ROI annotation objects created in section 1.3.
3.1.7 Update the name of nuclear stain channel in row 23.
3.1.8 Run the script by clicking on [Run] or by pressing Ctrl + r. The script is finished when an Info box is printed in the editor “Detected X number of cells” and the bottom of the editor displays “Stopped X:X:X”, as shown in Figure 19 below.

3.1.9 You should now be able to see cell detection objects within your annotation ROI in the image viewer and a list of the objects in the [Hierarchy] tab in the analysis menu (you may have to click on the arrow by the annotation ROI).
3.1.10 By clicking on [Measure] → [Show detection measurements] a table of all detection objects and their respective measurements will be shown. By clicking on [Show histograms] and choosing a measurement in the top pull-down menu, a summary of measurements can be seen for the cells.
3.1.11 To toggle detection overlays on and off, click on the [Show/hide detection objects] symbol, 
3.1.12 Optionally tweak the parameters in your script for a better segmentation, as described below, until you are satisfied with the results.
(row 36): value used to approximate cell bodies based on distance-based nucleus expansion. A greater value results in larger cell bodies. The parameter should be set according to the cell types present in the sample. Cell expansion is not considered in the later stages of the analysis workflow, and the value can be left as is for now.
(row 37): value used to constrain the cell expansion to a multiple of the nucleus size. A greater value results in a less strict cell expansion constraint and generally larger cell bodies. Similarly to cell expansion, cell constrain scale will not be considered at later stages and can be left as is for now.
(row 54): value used to set the minimum allowed nuclear area in µm2 for a cell. Smaller cells will be removed. The parameter should be set according to the cell types present in the sample.
(row 55): value used to set the minimum allowed mean nuclear intensity for a cell. Cells of lower intensities will be removed. The value should be set according to the nuclear staining intensities.
(rows 27-30): used to calculate preprocessing normalization based on the entire image to avoid detecting cells in negative regions.
-
: used to downsample the image for normalization. A greater value results in higher run times.
-
: used to normalize the image. Increasing the first value means that more of the darkest pixels are excluded, and increasing the second value means that more of the brightest pixels are excluded. Increase the values if you have artefacts, outliers or contrast issues in your image. Start with a value of 0.1, 99.9 and fine tune if needed.
(row 32): determines the minimum probability that a cell should have to be detected. Lower values may lead to over-detection of nuclei, while higher values may result in under-detection. A value of 0.5 is generally a good starting point.
(row 34): resolution of detection. Start with the same value as your voxel size and fine tune it if you want to better detect smaller or larger cells.
Note: Section 3.2 assigns signals to cells based on the distance from the nucleus boarder to the signal coordinate. Therefore, tuning cell expansion and cell constrain parameters won’t affect the signal mapping. However, due to a design limitation in QuPath, if cell segmentation is run without cell expansion, the resulting objects will not be of the cell object type. Since this limits the ability to identify cells at later steps, it is simplest to perform cell expansion either way but only regard the nuclei in the remaining analysis. Explanations for mapping signals based on the whole cell body is given within section 3.2 if desired.
3.1.13 (Optional) To run cell detection for multiple images in a project, click on [] → [Run for Project], mark the images in the “Available” box that you want to analyse and click on [>] to move them to the “Selected” Box. Once all images have been moved to the correct box, click on [OK]. A bar will now show to indicate how many of the images that are done. Once done, a pop-up window will show up asking if you want to reload the image you have opened, press [OK].
3.2 Signal mapping
Signal mapping is done to assign signals as child objects to cells, which later will be used for classifying cells as positive or negative for each marker. In this guide, a distance-based approach is applied where the cell with the shortest distance from its nucleus boundary to the signal is chosen as a parent. A threshold is chosen to set the maximum allowed distance that a signal can have from a nucleus boundary to be assigned as a child object.
3.2.1 Open the script editor and click on [File] → [New] or press Ctrl + N to open a new script. Copy and paste the code below. The script maps signals to cells by assigning them as cell child objects. Additionally, the script adds a measurement of the number of assigned signals of each signal class to the cell.
3.2.2 Change the distance parameter at row 21 to change the maximum allowed distance between a signal coordinate to the boundary of a cell. The signal will only be assigned if the distance is equal to or smaller than the distance threshold.
Note: Signals are defined as a detection object that is not of the object class “cell” (row 20). This needs to be redefined if there are other non “cell” detection objects that should not be sorted, for example by finding objects by classification:
3.2.3 Run the code by clicking on [Run] or Ctrl + R.
Note: QuPath works with objects in a hierarchical system with the image as the root-object. The system makes it easier to organize the analysis results and to call for detection objects within certain annotation ROIs. The hierarchies can be seen in the [Hierarchy] tab and the child objects are listed by clicking on the arrows on the left side of the parent object. More information regarding hierarchies can be found at QuPath’s documentation site[4].
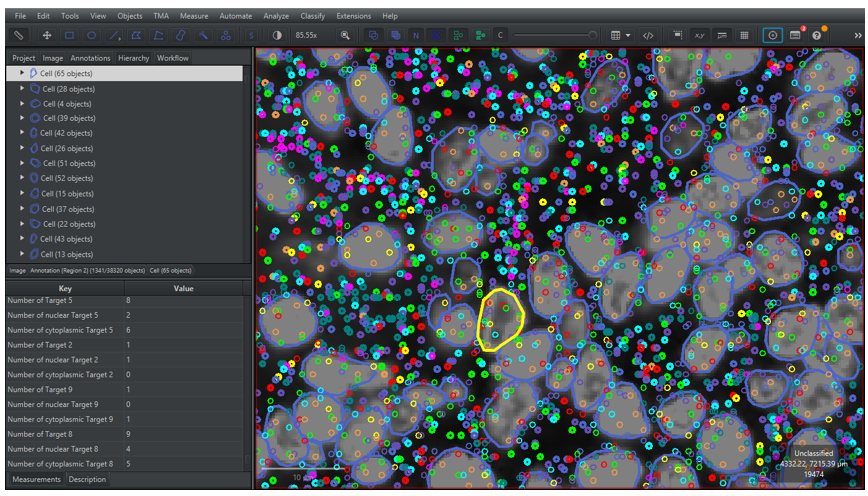
3.2.4 Go to the [Hierarchy] tab in the analysis menu to see the hierarchy of annotation and detection objects in the analysis panel. By clicking on the arrows on the left side of the cells, the signal child objects will be shown. Moreover, by selecting a cell the number of assigned signal object for each class can be seen in the measurement list as “Number of (component)(target))”, as shown in Figure 20 below. There are three different compartments:
- Nuclear: Signals overlap with the nuclear segmentation.
- Cytoplasmic: Signals do not overlap with the nuclear segmentation.
- If no compartment is specified, then it is a count for all signals assigned to the cell.
3.3 Cell classification
This section demonstrates how one can create cell classifiers in QuPath for multi-plex isPLA images. It should be noted that this is an example of how data created in section 2, 3.1 and 3.2 can be analysed, but many other methods exist to run cell classification.
3.3.1 Single measurement classifier
3.3.1.1 Go to the [Annotations] tab in the analysis menu. Add new classes for cell positivity, e.g. “Target X positive” or “+ Target X”, for each one of your targets by right-clicking in the classification list to the right and click on [Add/Remove] → [Add Class]. Write the name of the new class and then click on [OK]. Repeat the step for all targets. Alternatively, classes can be created based on channels by right-clicking on the classification list → [Populate from image channels]. Note that the classification name should not be the same as used for the signal detection.
3.3.1.2 Click on [Classify] → [Object Classification] → [Create single measurement classifier]. A new dialogue window will now pop up.
3.3.1.3 Choose [Cells] as Object Filter. Leave the Channel filter as [No filter (allow all channels)]. In the [Measurement] roll down list, choose one of the measurements “Number of [Target X]” created in the previous section for one of the targets, as shown in Figure 21.
3.3.1.4 To facilitate choosing an appropriate threshold for cell classification, the signal detection classes can be toggled off, as described in step 2.5.6.
3.3.1.5 Turn off the other channels by clicking on the [Brightness/Contrast] symbol and ticking of the channels, as described in step 1.2.2. It can be beneficial to open a separate viewer to show different zoom levels at the same time by [View] → [Show mini viewer].
3.3.1.6 In the [Above threshold], select the class created for the target. In the [Below threshold] select [Unclassified] if the negative cells should not be classified.
3.3.1.7 Tick on [Live preview] and drag the [Threshold] pointer to change the minimum number of signals for cell positivity. Cells that have equal to or more signals than the selected threshold will be assigned the classification selected in the [Above threshold], the remaining cells will be classified as the class specified in the [Below threshold]. The main viewer will update the cell classifications by changing the overlay colour to the class colour according to the classification list in the analysis pane, as shown in Figure 21. Alternatively set the number of signals that should facilitate a positive cell in the box next to the threshold pointer.
3.3.1.8 Select an appropriate threshold and write a name in [Classifier name] and press [Save]. A window will appear in the left down corner of the screen to inform that the classifier has been saved. All classifiers will automatically be saved in a folder named “classifiers” in the project folder.

3.3.1.9 Repeat steps 3.3.1.2 – 3.3.1.7 for each one of your targets.
Note: Using a machine learning model for cell classification can improve and advance your results beyond simple counts. QuPath has more information on how to do this in their documents.
3.3.2 Creating and running a composite classifier
In Section 3.3.1, classifiers were built to independently classify cells as positive or negative for each target. To identify cells that are positive for multiple targets simultaneously (e.g. cells positive for Target 1, Target 2 and Target 3), these individual classifiers are combined into one. The composite classifier will create new classes for all possible combinations of the classes and classify the cells accordingly.
3.3.2.1 Click on [Classify] → [Object Classification] → [Create composite classifier]. A new dialogue window will now pop up.
3.3.2.2 Select the classifiers created in section 3.3.1 above in the [Available] box and then click on the [>] to move them to [Selected], as shown in Figure 22. Enter a name for the new classifier in the [Classifier name] and click on [Save].
3.3.2.3 Apply the composite classifier by clicking on [Classify] → [Object Classification] → [Load object classifier], select the classifier and click on [Apply classifier]. Alternatively, copy the code below and replace the string to the name of your classifier.

Note: Objects in QuPath can only have a single classification. When applying multiple classifiers, the individual classes are pieced together into a composite class. For example, a cell classified as “+ Target 1”, “+ Target 2” and “+ Target 3” will be assigned the combined class “+ Target 1 : + Target 2 : + Target 3”.
3.3.2.4 (Optional) Add the new classes by clicking on the [Annotations] tab and right-clicking on the Classification list box → [Populate from existing objects] → [All classes]. This facilitates toggling the viewing of the objects on and off.
3.3.2.5 A summary of cell positivity can be seen by clicking on the annotation region of interest and looking at the measurement table on the lower left side.
3.3.2.6 Save the changes to the project when you are done.
3.4 Reviewing results
It is advisable to review the data output before continuing to downstream analysis. The following steps in this section can be performed for data review.
A simple co-occurrence matrix for marker positivity can be done by the following instructions:
3.4.1 Copy and paste the code below to a new script to print out a co-occurrence matrix for each main cell class (e.g. +Target 1, +Target 2 etc), as exemplified in Figure 23.
3.4.2 Add more targets to the cell classification list by removing the 

3.4.3 The script will be finished when the terminal has printed a header with “Co-occurrence matrix for ROIs [list with specified ROIs]” followed by a co-occurrence matrix. Copy the results that are printed in the terminal and paste it an empty excel sheet.
3.4.4 In Excel, select all numerical elements, go to [Home] -> [Conditional Formatting] -> [Colour scales] and choose one of the options. Now you can see an overview of all the co-occurrences of target positivity for the cells in the selected ROIs.
A bar graph to show the number of positive cells for each marker can be created in QuPath by the following instructions:
3.4.5 Copy and paste the code below to a new script. The script prints out bar graphs for each ROI class of the number of cells for each classification.
3.4.6 Add more targets to the cell classification list by removing the 


3.4.7 Run the script.
3.4.8 A dialogue window will appear with a bar-graph showing the fractions of cell positivity in each ROI, as shown in Figure 16.

This guide has provided an example workflow for analysis of isPLA data, an introduction of how to use QuPath, recommendations for how to detect isPLA signals and examples of how to perform cell segmentation, signal mapping to cells and multi-target cell classifications. There are many more implementations not covered by this guideline, be sure to tailor your image analysis workflow to suit your specific research questions.
If you have any remaining questions, please do not hesitate to contact our support.
 in the active rows to the desired channel names. The code will name the channels by the order of the channel list, meaning that the first active row will name channel one (C1) and so on.
in the active rows to the desired channel names. The code will name the channels by the order of the channel list, meaning that the first active row will name channel one (C1) and so on. or press m, and then select the annotation by double clicking on it. The annotation is moved by pressing the cursor down within the annotation and then dragging it around.
or press m, and then select the annotation by double clicking on it. The annotation is moved by pressing the cursor down within the annotation and then dragging it around. (row 36): value used to approximate cell bodies based on distance-based nucleus expansion. A greater value results in larger cell bodies. The parameter should be set according to the cell types present in the sample. C
(row 36): value used to approximate cell bodies based on distance-based nucleus expansion. A greater value results in larger cell bodies. The parameter should be set according to the cell types present in the sample. C (row 37): value used to constrain the cell expansion to a multiple of the nucleus size. A greater value results in a less strict cell expansion constraint and generally larger cell bodies. Similarly to cell expansion, cell constrain scale will not be considered at later stages and can be left as is for now.
(row 37): value used to constrain the cell expansion to a multiple of the nucleus size. A greater value results in a less strict cell expansion constraint and generally larger cell bodies. Similarly to cell expansion, cell constrain scale will not be considered at later stages and can be left as is for now.  (row 54): value used to set the minimum allowed nuclear area in µm
(row 54): value used to set the minimum allowed nuclear area in µm (row 55): value used to set the minimum allowed mean nuclear intensity for a cell. Cells of lower intensities will be removed. The value should be set according to the nuclear staining intensities.
(row 55): value used to set the minimum allowed mean nuclear intensity for a cell. Cells of lower intensities will be removed. The value should be set according to the nuclear staining intensities. (rows 27-30): used to calculate preprocessing normalization based on the entire image to avoid detecting cells in negative regions.
(rows 27-30): used to calculate preprocessing normalization based on the entire image to avoid detecting cells in negative regions.  : used to downsample the image for normalization. A greater value results in higher run times.
: used to downsample the image for normalization. A greater value results in higher run times.  : used to normalize the image. Increasing the first value means that more of the darkest pixels are excluded, and increasing the second value means that more of the brightest pixels are excluded. Increase the values if you have artefacts, outliers or contrast issues in your image. Start with a value of 0.1, 99.9 and fine tune if needed.
: used to normalize the image. Increasing the first value means that more of the darkest pixels are excluded, and increasing the second value means that more of the brightest pixels are excluded. Increase the values if you have artefacts, outliers or contrast issues in your image. Start with a value of 0.1, 99.9 and fine tune if needed.  (row 32): determines the minimum probability that a cell should have to be detected. Lower values may lead to over-detection of nuclei, while higher values may result in under-detection. A value of 0.5 is generally a good starting point.
(row 32): determines the minimum probability that a cell should have to be detected. Lower values may lead to over-detection of nuclei, while higher values may result in under-detection. A value of 0.5 is generally a good starting point.