This guide aims to support customers using the Omni kit in registering and merging images. The guide includes instructions for image and experimental requirements, installation, pipeline initiation, parameter selection, evaluation and lastly parameter optimization. Please make sure to read the pipeline requirements before running any experiments.
Guideline for image registration
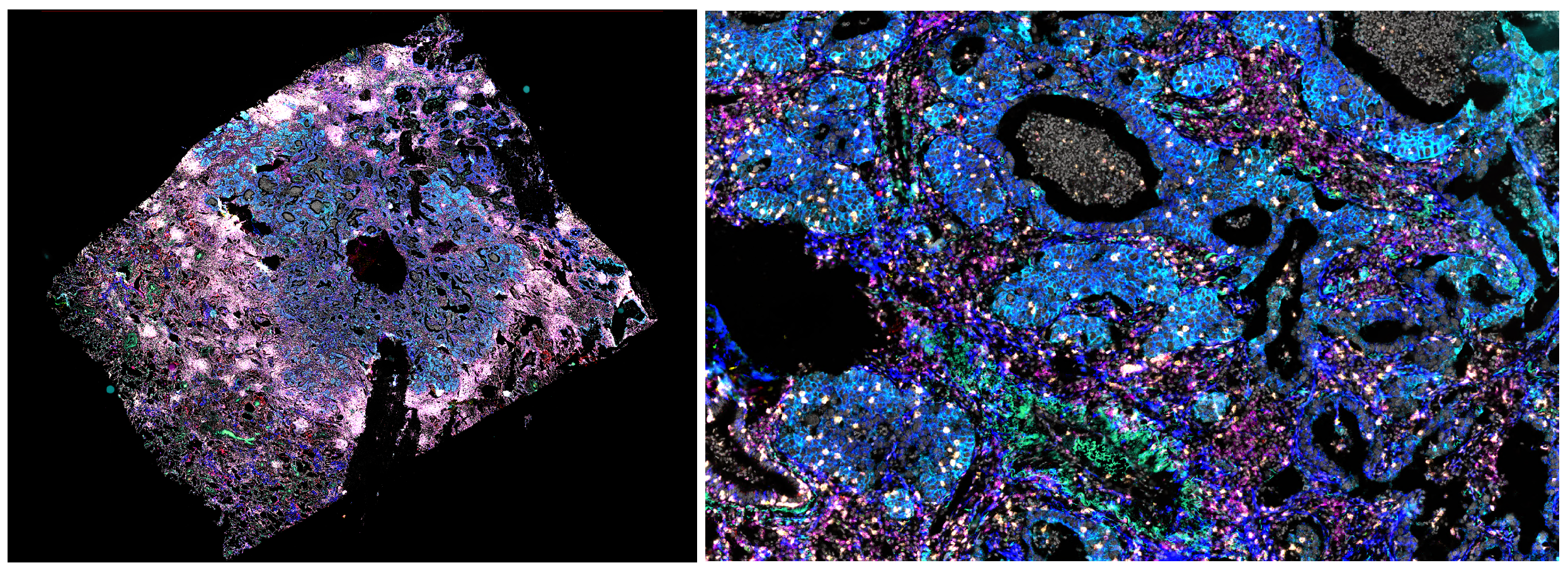
Image registration is the process of aligning images acquired at different time points. It is recommended to perform registration to enable local analysis, such as cell classification, of multiple isPLA stains from different cycles, and to facilitate background removal by subtracting autofluorescence channels acquired in a separate cycle. An example of an image produced with the Omni kit and registered with the workflow given in this instruction is shown in Figure 1 above. In short, image registration is performed by selecting a reference image, referred to as a fixed cycle, to which other images, referred to as moving cycle, are transformed. The transformation refers to a spatial mapping of points from the fixed image to the moving image, resulting in every pixel in the fixed image having a corresponding pixel in the moving image.
Contents
1 Pipeline requirements
2 Running the pipeline on Windows
2.1 Running the pipeline for the first time
2.2 Initiating the pipeline
2.3 Manual installation
3 Running the pipeline on mac
3.1 Running the pipeline for the first time
3.1.1 Installing Anaconda
3.1.2 Download Files
3.1.3 Setting up a virtual environment and installing Python packages
3.2 Starting the run on MacBook computers
4 Running the pipeline
5 Evaluating results
6 Fixing poor registration
7 Adapting the scripts
7.1 Changing file format naming
7.2 Adding additional transformations
8 Disclaimer
1. Pipeline requirements
The requirements specified in this section must be met to use the Registration pipeline in this guide.
Requirement 1: Two cycles are required
To use the registration pipeline, it is required to image at least two cycles of the same sample. The second cycle can be of other detection mix or of autofluorescence.
Requirement 2: All slides are stained and imaged similarly
It is important that the following is done the same between images and/or cycles that are registered simultaneously:
- All image regions have the same number of cycles
- All cycles have four channels: (1) one nuclear staining channel, (2) one channel detecting sets A, D or G, (3) one channel detecting sets B, E or H and (4) one channel detecting sets C, F or I. Autofluorescence or alternative cycles also require four channels.[1]
- The filter set order is the same between all images and cycles
Requirement 3: Images are in the correct file format
We recommend working with the following file formats:
- .ome.tif
- .ome.tiff
- .tif
- .tiff
- .btf
The following formats should be compatible with the registration pipeline, but have not yet been fully validated:
- .scn
- .ndpi
- .svs
- .czi
Requirement 4: Image cycles are saved in separate folders
Images acquired for different cycles must be saved in separate folders in the same directory. See an example of file organization at the bottom of this section.
In the examples in Section 4, a main folder ‘Project folder’ has been created in which the subfolders ‘Cycle-1_ABC’, ‘Cycle-2_DEF, ‘Cycle-3_GHI’ and ‘Cycle-4_AF’ sorts Images from the different cycles:
View Project Folder
Project folder /
Cycle-1_ABC /
Navinci_exp1_slide1_ch00.tif
Navinci_exp1_slide1_ch01.tif
Navinci_exp1_slide1_ch02.tif
Navinci_exp1_slide1_ch03.tif
Navinci_exp1_slide2_ch00.tif
…
Cycle-2_DEF /
Navinci_exp1_slide1_ch00.tif
…
Cycle-3_GHI /
Navinci_exp1_slide1_ch00.tif
…
Cycle-4_AF /
Navinci_exp1_slide1_ch00.tif
…
Requirement 6: Image file names must follow the specified format
Image file names must be named according to:
[Image name]_ch[channel index]_tile[tile row index]-[tile column index].[suffix]
Where the different colors specify a format group and brackets specify a variable input. Where the different colors specify a format group and brackets specify a variable input. Channel and tile specifications are optional and should only be used if images are single-channel and/or require stitching. Check below for examples of correct and incorrect image names.
Note that the groups must appear in the exact order shown.
The format groups are:
- Image name: required group that refers to the name of the sample and/or image region. Images for different cycles and/or channels and tiles are sorted and connected by the image name. The registered and merged image will be named using the image name.
- The input can be written with all symbols except for a hyphen, ‘-‘.
- The sequences ‘_ch’ and ‘_tile’ cannot be written within the input.
- Channel index: optional group if channels are saved into separate, single-layered gray scale images. The group specifies which channel index the image is for.
- The group is initialized by ‘_ch’.
- The first channel must have index zero.
- The index must be written with numbers.
- The index can be written with zeros before the actual index number, for example index 1 can be written as ‘_ch1’, ‘_ch01’, ‘_ch001’ and so on.
- Tile row index and tile column index: optional group to specify the tile position for unstitched images. Please note that requirements for image stitching are noted at the end of this section.
- The group is initialized by ‘_tile’.
- The first row and column indexes must have index zero.
- The indexes must be written with numbers.
- The indexes can be written with zeros before the actual index number, for example the tile in row 1 and column 1 can be written as ‘_tile1-1’, ‘_tile01-01’, ‘_tile001-001’and so on.
- Suffix: required group for file extension, e.g. ‘.tif’.
Examples of correct file names:
‘Experiment1_Slide01.tif’
‘Experiment1_Slide01_tile00-00.tif’
‘Experiment1_Slide01_ch00.tif’
‘Experiment1_Slide01_ch00_tile00-00.tif’
Examples of incorrect file names:
‘Experiment1_Slide01 _tile00-00_ch00.tif’ – Channel and tile specifications are in the wrong order
‘Experiment1_Slide01ch00tile00-00.tif’ – channel and tile specifications are found by ‘_ch’ and ‘_tile’, resulting in ‘ch00tile00-00’ being misinterpreted as part of the image name.
Recommendation 1: Run the pipeline on data with the same panel
Only combine data from one panel per image registration to reduce the risk of mixing up assay names.
Recommendation 2: Image different samples and/or fragmented samples in separate regions
If there are multiple samples on a slide, or if a single sample has been severely fragmented, use several distinct imaging areas instead of one large scan. Smaller scanning areas greatly improve image registration quality.
Recommendation 3: Use a computer with high CPU and RAM
It is recommended to use a computer with high-performance CPU and more RAM. The computer might run out of memory during registration for large images if not enough RAM is available.
Minimal RAM specification: 16GB
Requirement for image stitching:
Stitching is supported for single layered, single channel and non-pyramidal, images of the following format:
- .tif
- .tiff
- .ome.tif
- .ome.tiff
- .btf
[1] Discrepancies between images can be handled by running the pipeline several times. Discrepancies between cycles requires custom scripts.
2. Running the pipeline on Windows
This section is for users who are running the pipeline on a Windows computer. Section 1.1 provides instructions on how to install the pipeline as an application and Section 2.2 provides instructions for how to initiate the pipeline. An alternative ‘manual’ installation is provided in Section 2.3 for more experienced Python users.
2.1 Running the pipeline for the first time
This section is for users who are using the pipeline on a Windows computer for the first time. Here, an installer is used to install the pipeline as an app. Please note that Windows 11 is required and has at least 1.2GB of free space.
2.1.1 Download the ‘misPLA_registration_installer’ by clicking on the GitHub link below
- Locate the installation file within your directory and start the installation by double clicking on it. Step through the installer until it is complete, make sure to agree to set up a shortcut on your desktop and start menu.
- When the installation has run, the application will be initiated automatically, and a terminal and dialogue window will appear on your screen, as shown in Figure 2. If nothing appears, look for the icon
in the taskbar and click on it. Skip ahead to Section 4.
2.2 Initiating the pipeline
- Initiate the application by double-clicking on the short-cut “misPLA registration” created on your desktop. Alternatively, search for “misPLA registration” in your Start Menu.
- If a short-cut was created on your desktop or homepage, then you can initiate the application by double clicking on it. Alternatively, locate the ‘misPLA registration.exe’ file on your computer. This is easiest done by clicking on ‘Search’ in your taskbar and typing in the name. Double-click on the file when you have located it to initialize it.
- A terminal and dialogue window should appear, as shown in Figure 2 below. If the pop-up window does not appear on the screen, look for the icon
in the taskbar and click on it.
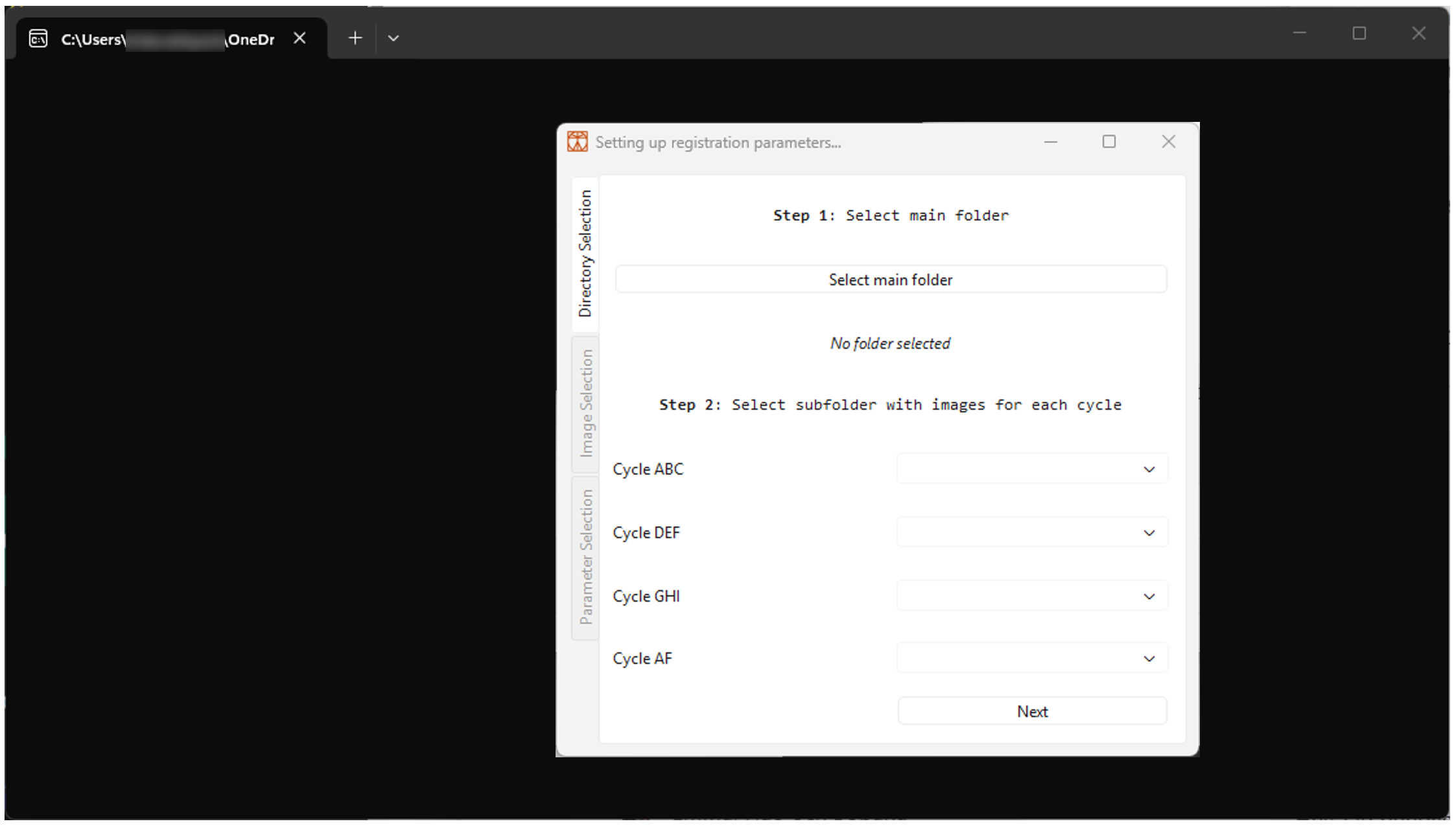
2.3 Manual installation
Manual installation requires Python to be installed on the computer with a version higher than 3.13.1.
2.3.1 To download the GUI application file, go to the GitHub Gist page linked below. Download both files by clicking on the download icon, .
2.3.2 Unzip the folder in your preferred location and rename it ‘misPLA_registration’.
2.3.3 To download the registration package, go to the GitHub Gist page linked below and download the files.
2.3.4 Unzip the folder within your ‘misPLA_registration’ folder and rename it “registration“.
2.3.5 Create a virtual environment ‘misPLA_registration.venv’ in your preferred location, activate it and install all modules specified in the ‘requirements.txt’ file downloaded with the GUI application file.
2.3.6 Initiate the pipeline by running the ‘GUI_registration.py’ file. A pop-up window should appear on the screen, as shown in Figure 2 above. If the pop-up window does not appear on the screen, look for the icon in the taskbar and click on it.
3. Running the pipeline on Mac
Running the pipeline as an executable application is not supported for Mac-users. Therefore, a manual set-up is required.
3.1 Running the pipeline for the first time
3.1.1 Installing Anaconda
Note, Miniconda can be installed as an alternative to Anaconda
Go to Anaconda (https://www.anaconda.com/download)
3.1.1.1 Download and install Anaconda
3.1.1.2 After installation, open a terminal and you should see (base) in the command line prompt. This indicates that you are in your base conda environment.
3.1.2 Download Files
3.1.2.1 To download the GUI application file, go to the GitHub Gist page linked below. Download both files by clicking on the download icon .
3.1.2.2 Unzip the folder in your preferred location and rename it ‘misPLA_registration’.
3.1.2.3 To download the registration package, go to the GitHub Gist page linked below and download the files.
3.1.2.4 Unzip the folder within your ‘misPLA_registration’ folder and rename it “registration”.
3.1.3 Setting up a virtual environment and installing Python packages
After installing Anaconda, create and set up a conda environment to run the pipeline
Create a dedicated conda environment by opening a terminal and running the command ‘conda create –n .ven python==3.13.1’. Note that python with a version higher than 3.13.1 is required.
3.1.3.1 Activate .ven conda environment using the command ‘conda activate .ven’
3.1.3.2 Install the required dependencies found in the file ‘requirements.txt’ downloaded with the GUI application file.
3.1.3.3 Locate to the ‘Customer-registration/registration’ folder, with the command ‘cd /Users/navinci/repos/Customer-registration/registration’. You should see a file called ‘requirements.txt’ under the folder.
3.1.3.4 Install the requirements by running the command ‘pip install –r requirements.txt’.
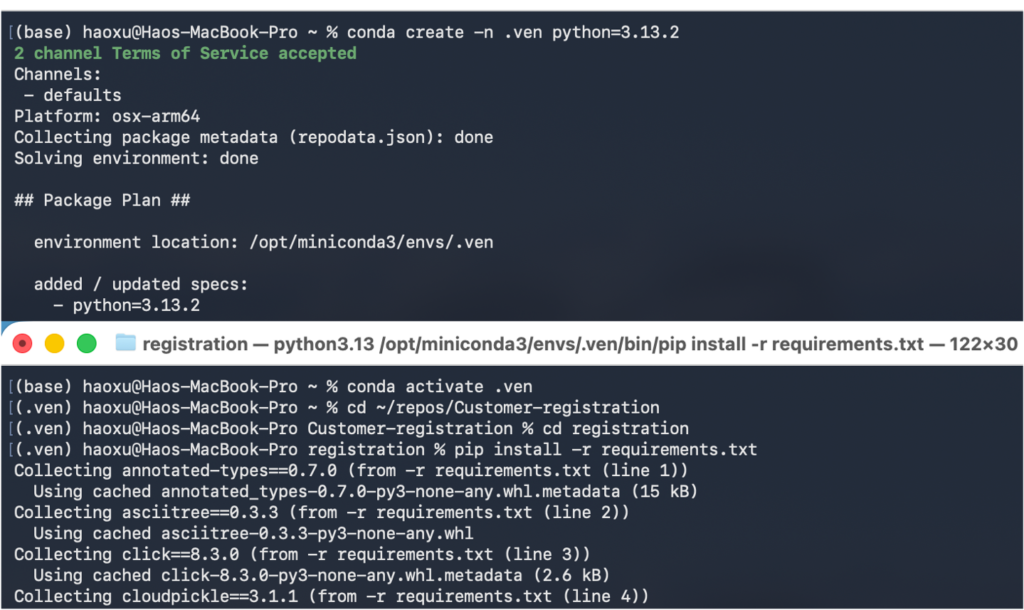
3.2 Starting the run on MacBook computers
Once all files and packages have been downloaded and installed, the script can be initiated.
3.2.1 Open a terminal and activate the conda environment by ‘conda activate .ven’
3.2.2 Locate the ‘Registration’ folder downloaded, e.g., ‘cd /Users/navinci/repos/Customer-registration’
3.2.3 Run the image registration script by command line ‘python GUI_registration.py’. A pop-up window should appear on the screen, as shown in Figure 4 below. If the pop-up window does not appear on the screen, look for the icon in the taskbar and click on it.
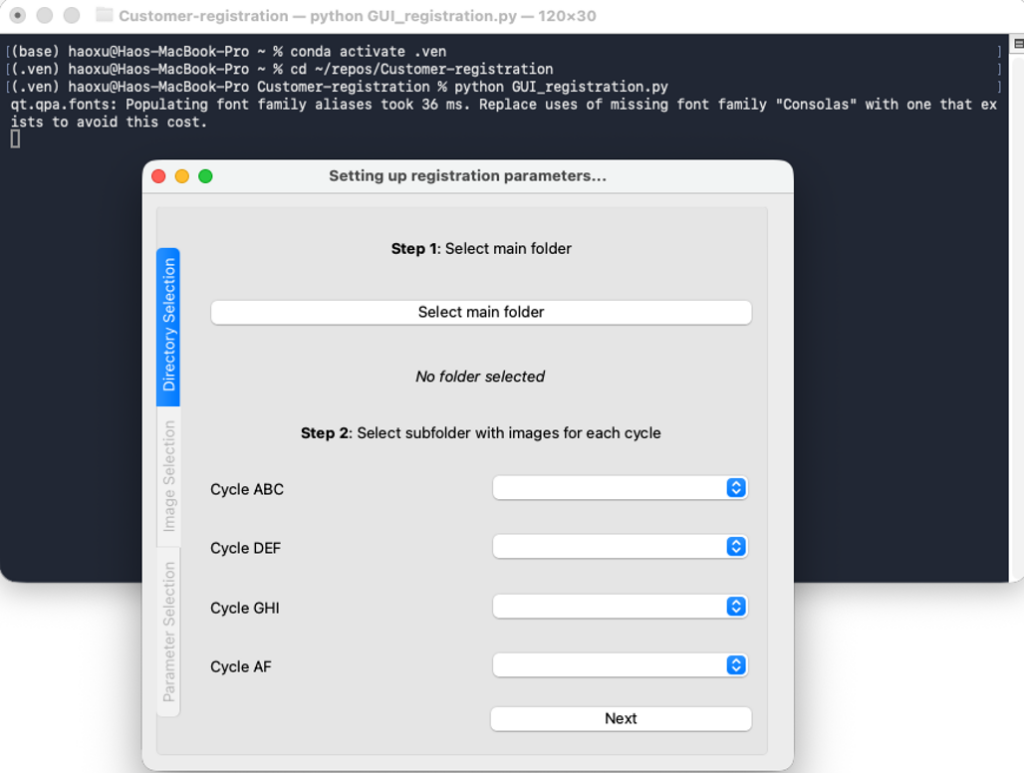
4. Running the pipeline
This section provides instructions on how to select parameters for image registration and describes what each parameter means. Please follow the instructions in Sections 2 and 3 before reading this section. Shortly after initiating the scripts as explained in Section 2.2 and 3.2, a pop-up window should appear as shown in Figure 2 (Windows) or Figure 4 (Mac) above.
4.1 Follow Step 1 in the Registration window by clicking on the button ‘Select main folder’. In the file explorer, locate the main folder containing the cycle sub folders with images and click on ‘Select Folder’.
4.2 In the registration window, follow step 2 and select subfolders for each cycle, as shown in Figure 5 below. The cycle refers to the detection mix imaged, for example ‘Cycle ABC’ refers to imaging of ‘Cycle 1/ABC’ instructed in the ‘Instructions for Use’ of the Omni kit.
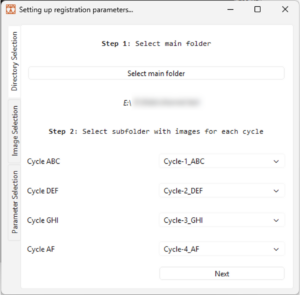
4.3 Click on the ‘Next’ button or on the ‘Image Selection’ in the menu to the left.
4.4 Follow step 3 by choosing slides to register.
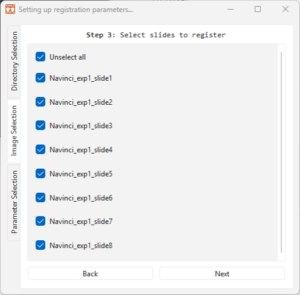
4.5 Click on the ‘Next’ button or on the ‘Parameter Selection’ in the menu to the right.
4.6 Follow step 4 by selecting the correct channel index for each filter set, starting from channel 0, as shown in Figure 7. The filter sets correspond to the filters specified in the Omni kit ‘Instructions for Use’ (table 4), where sets A, D, G are imaged with Cy5, sets B, E, H are imaged with Cy3 and sets C, F and I are imaged with FITC. If alternative filter sets have been used, then specify its corresponding filter set in the list.
4.7 Follow step 5 by selecting the general registration parameters, as shown in Figure 7 and as explained below. In short, select…
… the second cycle for the parameter Fixed cycle.
… Downsample factor 1 for grown cells and downsample factor 2 for tissue.
… Transformation ‘rigid, affine’ for grown cells and ‘rigid, affine, nl’ for tissue.
Fixed cycle: the fixed cycle refers to the cycle that is fixed and non-transformed during registration. With other words, all other cycles are transformed and registered based on the fixed cycle. Choose the middle time point cycle to have as little morphological change between the fixed and moving cycles. For example, if cycles ABC, DEF and GHI have been imaged in order, then select cycle DEF. If there is an even number of cycles, select the one at an earlier time point. For example, if cycles ABC, DEF, GHI and AF (autofluorescence) has been imaged in that order, then select cycle DEF. If the selected fixed cycle has obvious defects, it might be better to use another.
Downsample factor: the downsample factor refers to the degree of downsampling before calculating transformations for registration. A lower downsampling usually results in better results but a longer processing time.
It is recommended to choose factor 1 for grown cells and factor 2 for tissue when imaging with a 20X objective.
Transformations: the transformation parameter refers to how moving images are mapped to the fixed image:
Rigid – by rotation and translation
Affine – by shearing, scaling, rotation and translation
NL – with localized deformations (non-linear)
The transformations are added in sequence to add complexity to the registration sequentially. It is recommended to select ‘Rigid, affine’ for grown cells and ‘Rigid, affine, nl’ for tissue.
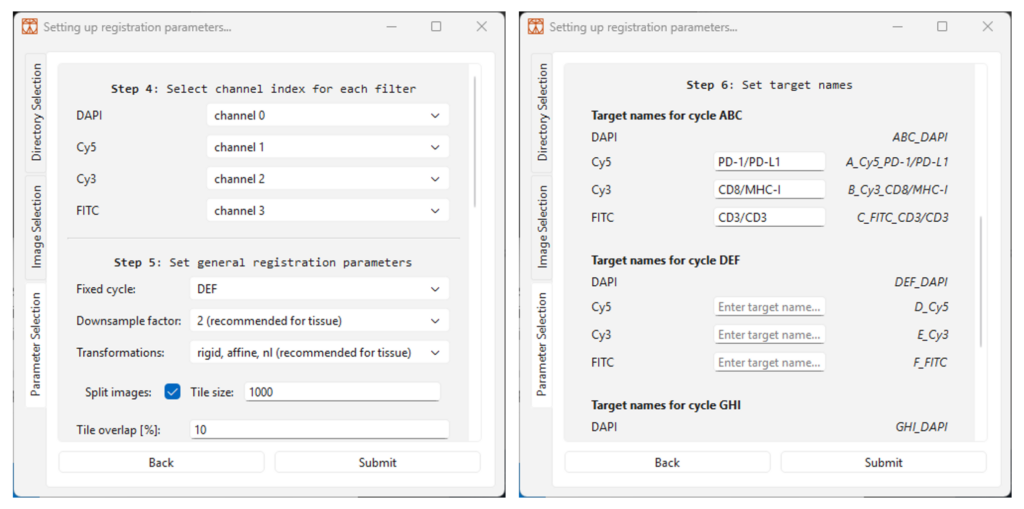
4.8 If your image names have a tile reference, an additional parameter “Tile overlap [%]”will automatically be added.
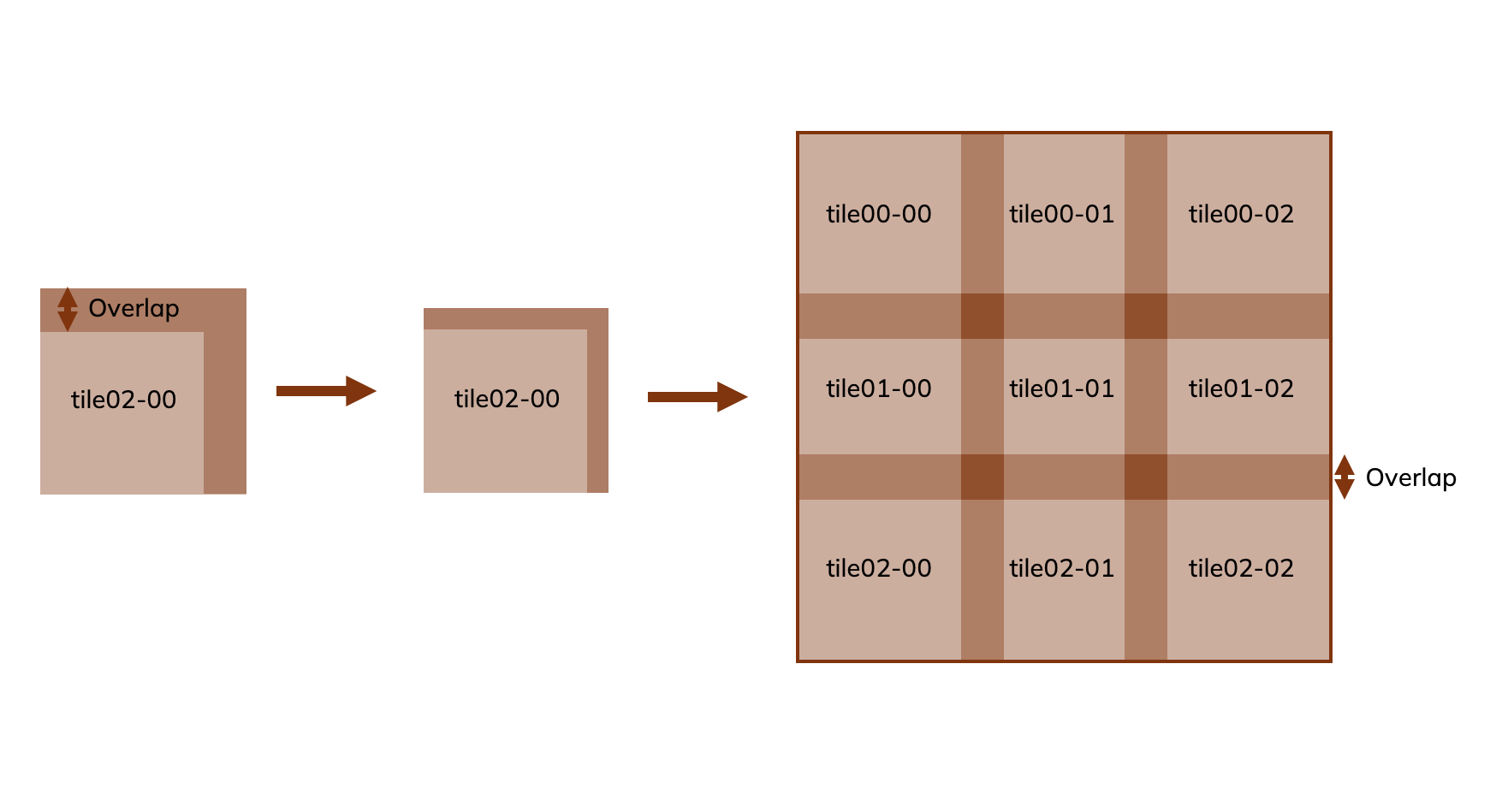
Tile overlap: The overlap between tiles in percentage.
The provided stitching script assembles tile images into a single mosaic based solely on the specified layout information, as shown in Figure 8 below. This step is intended as a simple pre‑processing stage before performing image registration. Note that the stitching method does not apply any affine transformations or geometric corrections—it only places tiles together, with or without overlap, exactly as given.
If you require stitching that incorporates affine transformations, you may find tools such as MIST in Fiji/ImageJ more suitable.
4.9 Follow step 6 by specifying image output parameters.
Compression: Select image compression type. Compression reduces file size but does not change image pixel values. All available methods are lossless. Select between deflate, lzma, zstd and packbits or select “none” to not compress the output images.
PackBits: Very fast compression. Provides limited file size reduction and works best for images with large uniform areas.
Deflate: A general purpose compression method that works well for many different image types. Good balance between file size and performance.
LZMA: Achieves high compression (small file sizes) but is slower when writing images.
Zstd: Achieves high compression (small file sizes) and fast reading and writing performance
MisPLA bit-depth: The bit depth for the multiplex isPLA image. Select “Original” to preserve bit depth and pixel values. Note that lowering the bit depth reduces the file size but may reduce precision in pixel values.
Reg ctrl bit-depth: The bit depth for the registration control isPLA image. Select “Original” to preserve bit depth and pixel values. Note that lowering the bit depth may reduce precision in pixel values.
MisPLA downsample factor: The downsample factor of the multiplex isPLA image. Downsampling is the process where several neighboring pixels are merged into one. As an example, a downsample factor of two produces an image of 1/2 of the original width and height in pixels, meaning each pixel represents an area that is four times larger than the original pixel area. A downsample factor of 1 keeps the original resolution.
Reg ctrl downsample factor: The downsample factor of the registration control image.
Split images: add an extra step of splitting images to tiles by checking the box. The output image (tile) size can be specified in the ‘Tile size’ parameter. Split images will be saved in a separate sub-folder. Please note that incomplete tiles, tiles smaller than the given tile size, will not be saved.
4.10 Follow step 7 by writing the target names for each filter set. As shown in Figure 7 above, the resulting channel name can be seen to the right of the editing box. It should be noted that DAPI channels, AF (Autofluorescence) cycles or additional cycles cannot be edited.
4.11 When all parameters are set, click on the ‘Submit’ button.
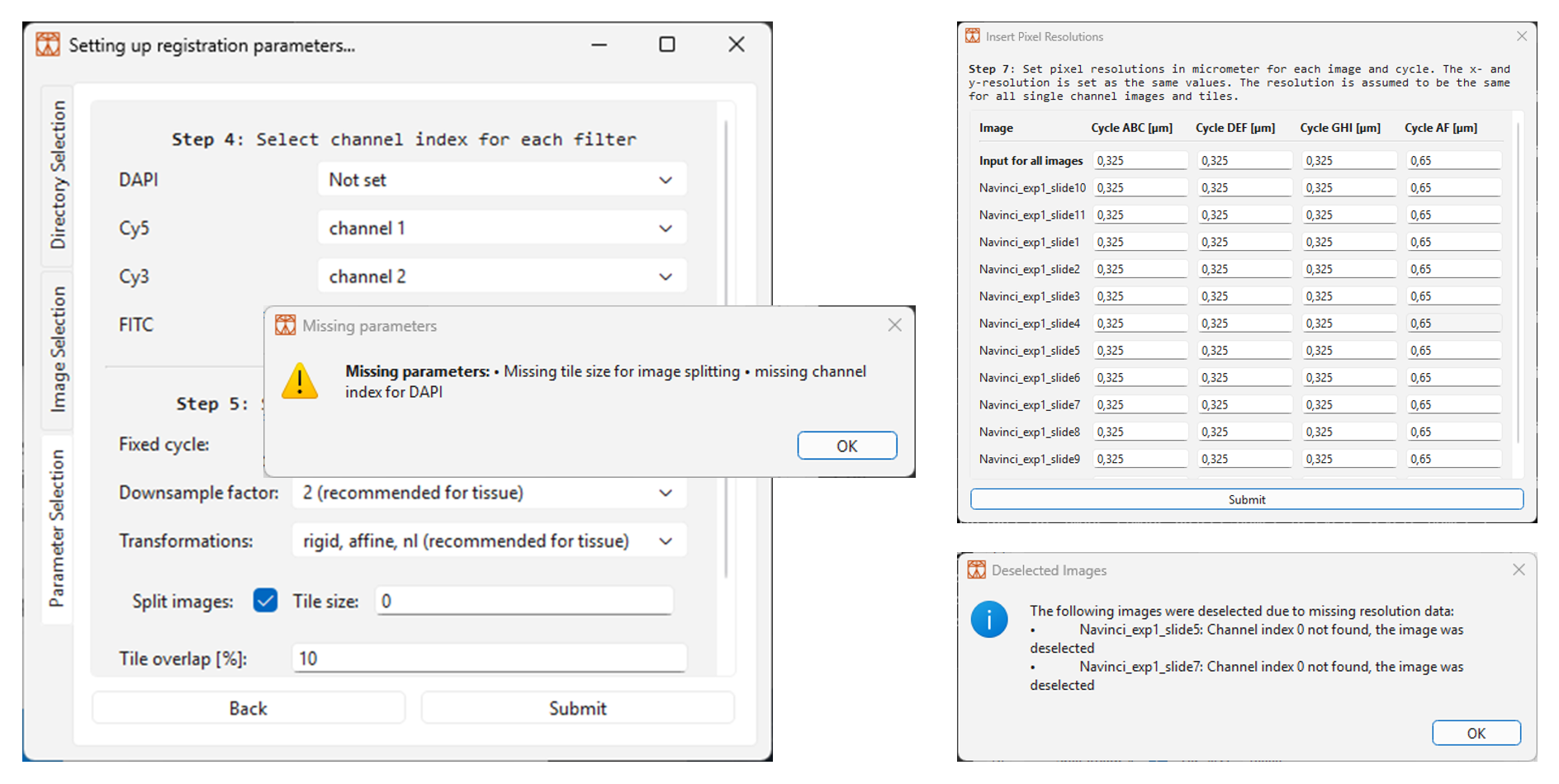
a. If a parameter is missing, a pop-up window will specify which ones need to be added, as shown in Figure 9 below.
b. If the software doesn’t succeed in reading pixel sizes, then a dialogue window will appear for user input, as shown in Figure 9 below. Add the pixel size in micrometer in the boxes and then click on ‘Submit’ to continue. Note that pixel width and height are assumed to be the same. Use the ‘Input for all images’ to set a value for all images.
c. If images are deselected due to missing data, an information dialogue window will appear. In the example in Figure 9 below, the images Navinci_exp1_slide5 and Navinci_exp1_slide7 have been deselected due to the pipeline not finding channel index zero. Check the terminal for more information.
When all parameters have been submitted, the terminal and dialogue window will show the registration progress, as shown in Figure 10 below. Note that any errors occurring in the script will be written in the dialogue window and that the script will continue running for the remaining images.
For an image of size ~2 cm2, each image takes around 50 minutes. This is a rough estimate and depends on the computer as well as the number of cycles and image size.
4.12 Once the script has finished running for all images, look through the text in the dialogue window to see if everything has been run correctly. An error will be printed in the dialogue window If something has failed. Click on ‘Close Window’ to shut down the terminal and dialogue window.
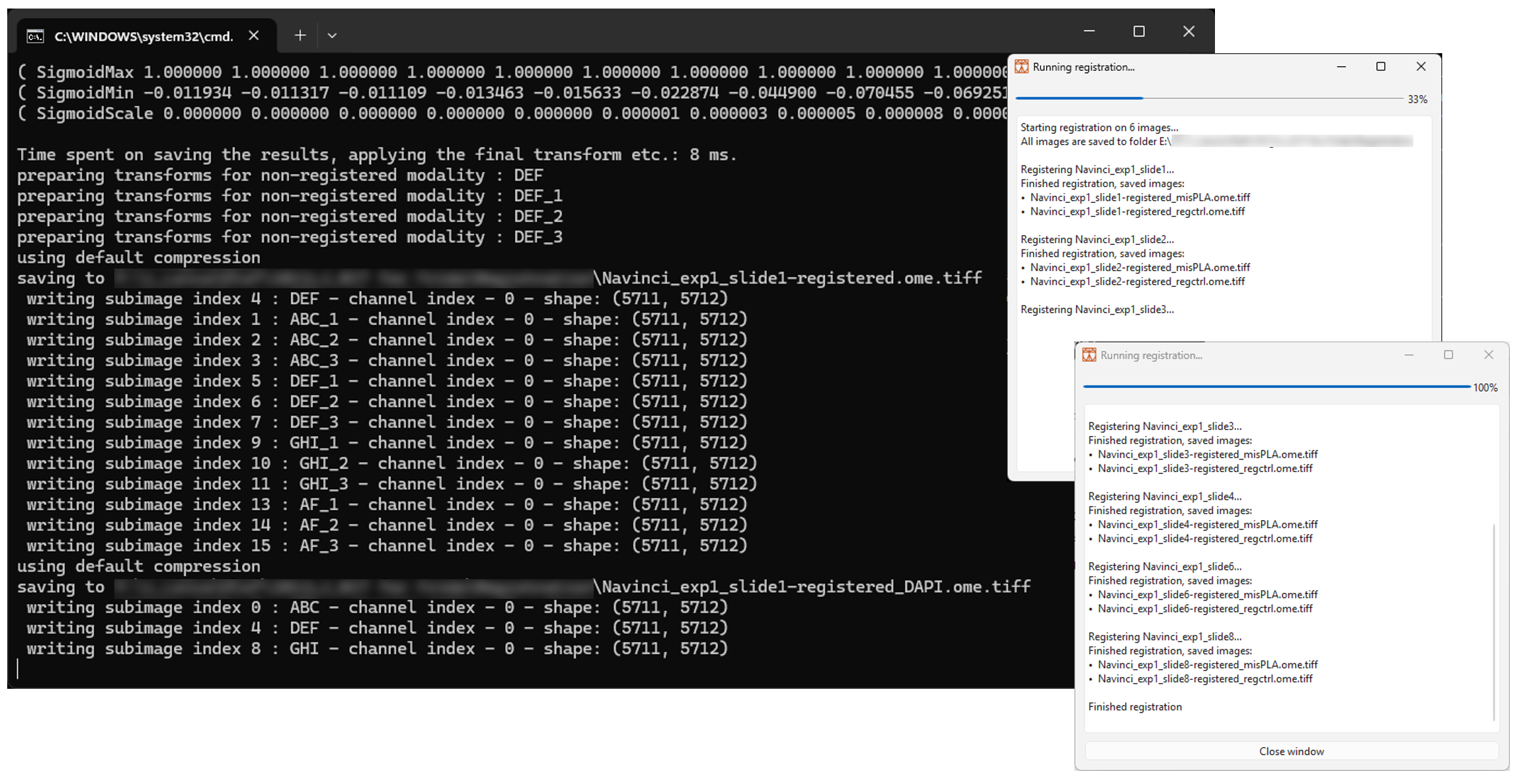
Multiple folders and files have been created in the subfolder ‘Registration’ in the project folder:
- Registered images: ‘[image]-registered_misPLA.ome.tiff’ files are images containing the DAPI channel from the fixed cycle and all cycles’ isPLA channels.
- Registration control images: ‘[image]-registered_regctrl.ome.tiff’ files are images containing all cycle’s DAPI channels.
- ‘transformations’: a folder containing transformation files for each registration. There is one subfolder per slide and moving cycle, all folder names are written accordingly: ‘[image]-[moved cycle]_to_[fixed cycle]_reg_output’.
- ‘configurations’: ‘.yaml’ configuration set-up files containing parameters and maps for all slides registered
- ‘Stitched’: a folder containing the stitched images before transformation and registration, if stitching has been done. Stitched images will be saved in cycle sub folders.
- ‘Split images’: a folder containing tiled images, if the function ‘Split images’ have been selected in step 4.9. Each registered image will have its own separate sub folder, and each tiled image will be named ‘[registered image name]_tile[index]_[x start position]_[y start position].ome.tiff”.
See section 5 for instructions on how to evaluate the results.
5. Evaluating results
5.1 Open your preferred image visualization software, QuPath is used in the examples below.
a. Note that the software should be compatible with ‘.ome.tiff’ images and with the file size of your output images.
b. For first-time QuPath users, please check out the QuPath guideline for instructions on how to open images.
5.2 Upload the Registration misPLA and Registration control images as instructed for the software. To evaluate the results, open the Registration control images one at a time.
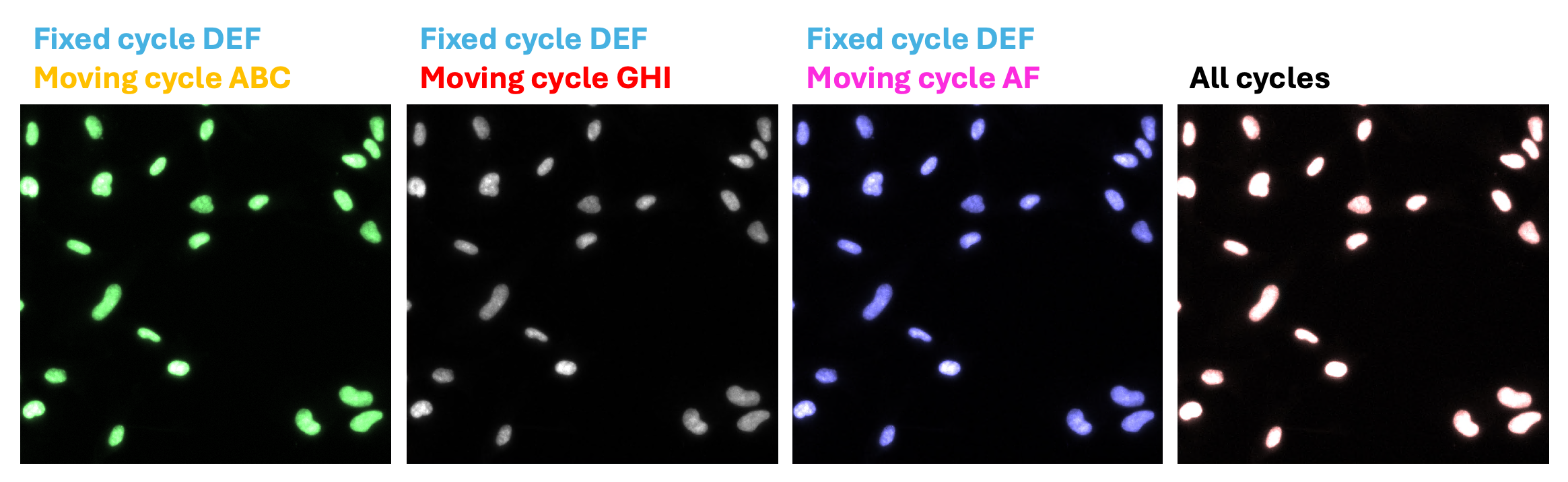
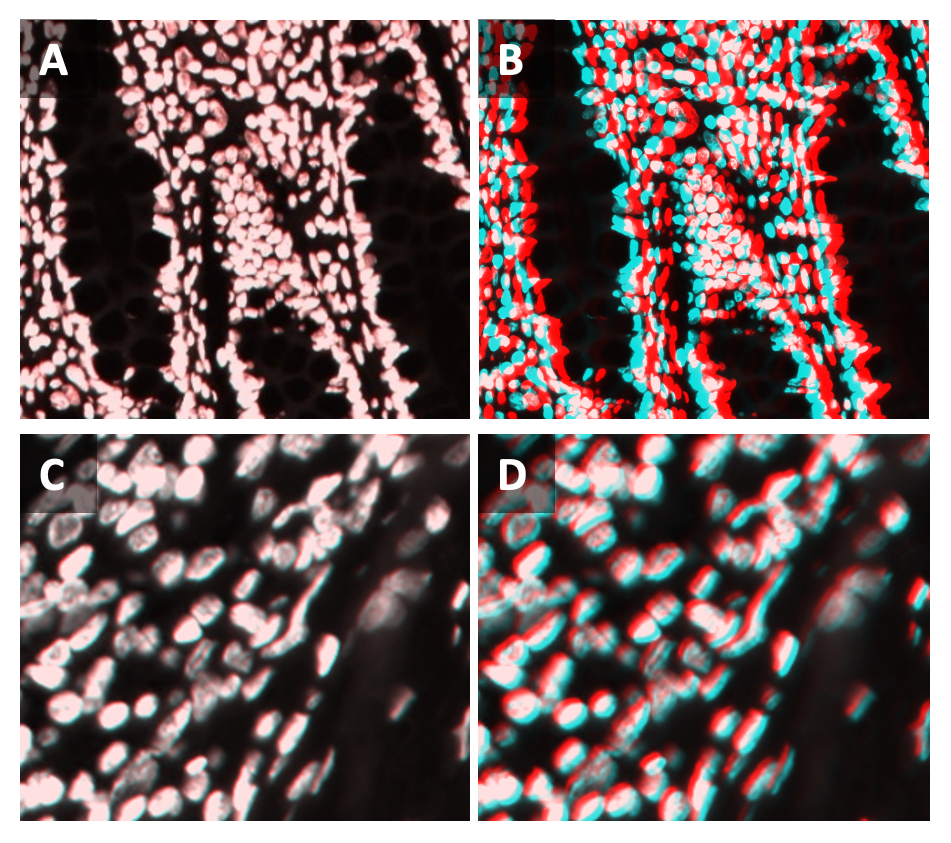
5.3 Either evaluate the images by looking at all DAPI channels together or by looking at the fixed cycle with the moving cycles one at a time. It is recommended to visualize the channels with complementary colors, as shown in Figure 11 below:
Fixed channel: Cyan
Moving channel: Yellow → green color indicates good alignment
Moving channel: Red → grey color indicates good alignment
Moving channel: Magenta → purple color indicates good alignment
All channels together: white indicates good alignment between all cycles

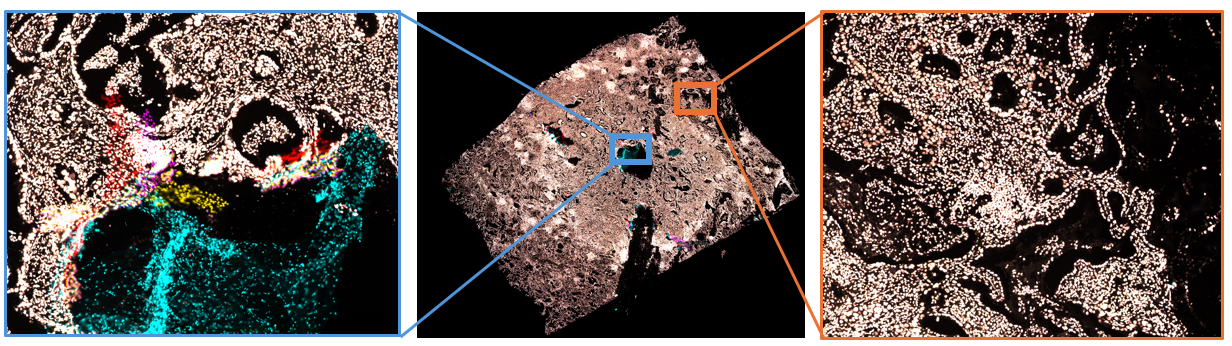
Note that poor alignment is expected around regions with artefacts and morphological changes, as shown in Figure 12 below. Artefacts that can cause misalignment include:
- Poor tile stitching
- Tissue folds and tears created between cycles
- Local movement of grown cells or regions between cycles
- Staining smears and artefacts in the nuclear staining channel
- Poor focus
- Mounting media bubbles

If the registration control images show overall good nuclei alignment, continue with analysis and exclude regions of poor alignment. Examples of acceptable and poor alignment are shown in Figure 13 below. If the images show poor misalignment, continue with section 6 to optimize parameters and improve registration results.
6. Fixing poor registration
Important notes before re-running the registration pipeline:
- Images with acceptable alignment can be excluded in the ‘Image Selection’ tab, as explained in step 4.4.
- Re-running the registration of an image will overwrite the old output files. To save old files, make sure to move them to another location or rename them.
- If your images have been stitched within the registration pipeline and you are pleased with the result, you should select the ‘Stitched’ subfolder as project folder to avoid redoing the stitching. This will however save all new files and folders within the new project folder, and old results will be kept in the previously selected folder.
Before testing registration with different parameters, it should be noted that the image registration will be different every time it is run for an image, meaning that faulty registration can sometimes be fixed by simply re-running the pipeline.
If re-running the pipeline did not improve the registration, then please test the following:
- Try registration with a lower downsample factor (if not already set to 1). A lower downsample factor generally results in better registration results. It should be noted that a lower factor requires higher computational capacity and might run for a longer period of time.
- Try registration with simpler transformations:
- For registration of tissue where the transformations ‘rigid, affine, nl’ have been used, try using ‘rigid, affine’ instead. Removing the non-linear transformation should improve overall alignment, although local deformations will not be accounted for.
- If transformations ‘rigid, affine’ have been used, try using ‘rigid’ instead.
- Try cropping and/or splitting up your images.
7. Adapting the scripts
This section provides instructions on how to adapt the script for specific data sets, for example when the file format names do not match the requirements listed in section 1. Caution is advised when modifying the script as such changes may impact its overall functionality.
7.1 Changing file format naming
For users who have downloaded the pipeline as a Windows application, as instructed in Section 2.1, the registration module scripts can be found within the downloaded folder ‘/_internal/registration’.
To change the file format naming, the parameter file_name_pattern at line 409 in ‘registration.reg_classes.py’ must be changed. The following must be met when modifying the parameter:
- Groups must be written in the same order as the file name.
- Each group is written as ([group name], [literal], [regex pattern], [required]):
- Group name: the name of the group, which can be either “name”, “channel” or “tile”
- Literal: defines if a literal is used before the group as a separator. If no literal is used, then use an empty string, “”.
- Regex pattern: A regular expression pattern specifying how the group is identified
- Required: Boolean value to set if the group is required (‘True’) or optional (‘False’).
Please note that adding groups might require adapting other parts of the script.
As an example, the requirements specified in section 1 results in the following groups:
- Image name:
- Group name: “name”
- Literal: “”, referring to no literal specified before the group
- Regex pattern: “(?:(?!ch|tile).)+?”, which asserts that the groups does not contain the strings ‘ch’ or ‘tile’. All other characters are allowed at any length.
- Required: True, meaning that there must be a match
- Channel:
- Group name: “channel”
- Literal: “_”, meaning that ‘_’ must be added before the group
- Regex pattern: r”ch\d{2}”, which asserts that the group is initiated by ‘ch’ followed by two digits.
- Required: False, meaning that the group is optional
- Channel:
- Group name: “tile”
- Literal: “_”, meaning that ‘_’ must be added before the group
- Regex pattern: r”tile\d{2}-\d{2}”, which asserts that the group is initiated by ‘tile’, followed by two digits, a ‘-’ and two digits.
- , which asserts that the group is initiated by ‘ch’ followed by two numbers.
- Required: False, meaning that the group is optional
7.2 Adding additional transformations
Additional registration parameters can be added as options in line 344 in ‘reg_classes.py’ found in the subfolder ‘registration’. Multiple parameters are written as a list in the same order as they should be applied. The parameters should be written as strings and must be listed in wsireg.parameter_maps.reg_params, for example:
- ‘similarity’
- ‘fi_correction’
- ‘nl-reduced’
- ‘nl-mid’
- ‘rigid-expanded’
- ‘nl3’
- ‘nl2’
For example, [“rigid”,”affine”,”nl”], as written in line 348 in ‘reg_classes.py’ refers to parameter maps ‘rigid’, ‘affine’ and ‘nl’ being used sequentially.
8. Disclaimer
This material is provided solely for Navinci product analysis. Any other use, disclosure, or distribution is prohibited unless authorized. The material is provided “as is,” without warranties.